
| Sequence ID | dm3.chrX |
|---|---|
| Location | 13,588,915 – 13,589,080 |
| Length | 165 |
| Max. P | 0.884813 |

| Location | 13,588,915 – 13,589,014 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 81.26 |
| Shannon entropy | 0.33727 |
| G+C content | 0.42247 |
| Mean single sequence MFE | -26.80 |
| Consensus MFE | -15.52 |
| Energy contribution | -16.24 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.07 |
| SVM RNA-class probability | 0.884813 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
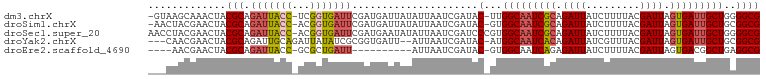
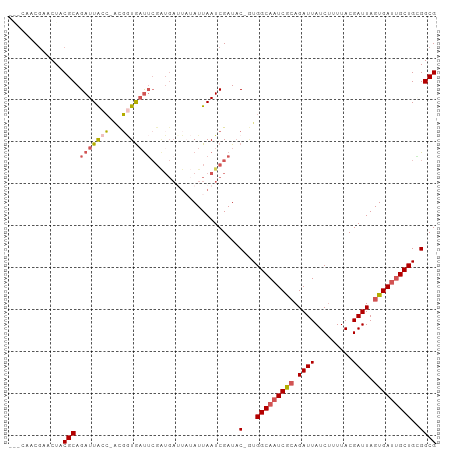
>dm3.chrX 13588915 99 + 22422827 -GUAAGCAAACUACGCAGAUUACC-UCGGUGAUUCGAUGAUUAUAUUAAUCGAUAC-UUGGCAAUCGCAGAUUAUCUUUUACGAUUAGUGAUUGCUGGGGCG -............(((..(((((.-...)))))(((((..........)))))..(-(..((((((((.((((.........)))).))))))))..))))) ( -27.10, z-score = -2.37, R) >droSim1.chrX 10416969 99 + 17042790 -AACUACGAACUACGCAGAUUACC-ACGGUGAUUCGAUGAUUAUAUUAAUCGAUAC-GUGGCAAUCGCAGAUUAUCUUUUACGAUUAGUGAUUGCUGCGGCG -............(((..(((((.-...)))))(((((..........)))))..(-(..((((((((.((((.........)))).))))))))..))))) ( -28.30, z-score = -2.76, R) >droSec1.super_20 227513 101 + 1148123 AACCUACGAACUACGCAGAUUACC-ACGGUGAUUCGAUGAAUAUAUUAAUCGAUCCCGUGGCAAUCGCAGAUUAUCUUUUACGAUUAGUGAUUGCUGGGGCG ......((..(((.(((((((.((-((((.(((.((((..........))))))))))))).))))............((((.....)))).))))))..)) ( -29.90, z-score = -2.73, R) >droYak2.chrX 7872375 96 + 21770863 ---CAACGAACUACGCAGAUUGCAGAUUAUAUCGCGGUGAUU--AUUAAUCGAUAC-AUGGCAAUCACAGAUUAUCGUUUACGAUUAGUGAUUGCUGCGGCG ---...........((.....))........((((((..(((--((((((((.((.-(((..((((...))))..))).)))))))))))))..)))))).. ( -30.20, z-score = -3.27, R) >droEre2.scaffold_4690 5453552 86 - 18748788 ----AACGAACUACGCAGAUUACC-GCGCUGAUU----------AUUAAUCGAUAC-GUGGCAAUCAGAGAUUAUCUUUUACGAUUAGUGACGGCUGAGGCG ----.........(((...(((((-(((((((((----------.......(((..-......)))((((.....))))...)))))))).))).))).))) ( -18.50, z-score = -0.35, R) >consensus ___CAACGAACUACGCAGAUUACC_ACGGUGAUUCGAUGAUUAUAUUAAUCGAUAC_GUGGCAAUCGCAGAUUAUCUUUUACGAUUAGUGAUUGCUGCGGCG .............(((.(((((((...))))))).........................(((((((((.((((.........)))).)))))))))...))) (-15.52 = -16.24 + 0.72)

| Location | 13,588,978 – 13,589,080 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 80.06 |
| Shannon entropy | 0.34131 |
| G+C content | 0.41652 |
| Mean single sequence MFE | -12.76 |
| Consensus MFE | -10.80 |
| Energy contribution | -11.44 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.14 |
| Mean z-score | -0.88 |
| Structure conservation index | 0.85 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.592324 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
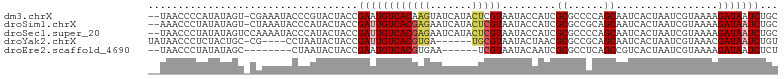
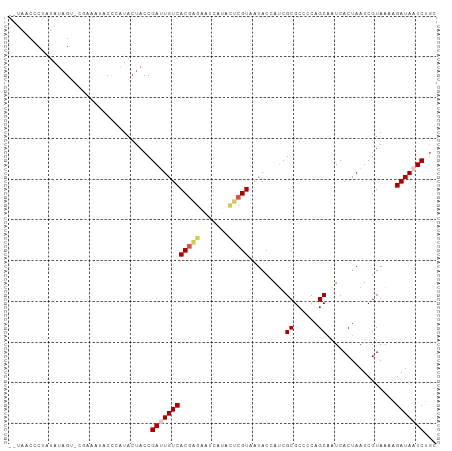
>dm3.chrX 13588978 102 - 22422827 --UAACCCCAUAUAGU-CGAAAUACCCGUACUACCGAAUGUCACAAGUAUCAUACUCGUAAUACCAUCGCGCCCCAGCAAUCACUAAUCGUAAAAGAUAAUCUGC --..........((((-((.......)).))))..((.((((((.((((...)))).)).........((......)).................)))).))... ( -10.00, z-score = -0.15, R) >droSim1.chrX 10417032 102 - 17042790 --AAACCCUAUAUAGU-CUAAAUACCCAUACUACCGAUUGUCACGAGAAUCAUACUCGUAAUACCAUCGCGCCGCAGCAAUCACUAAUCGUAAAAGAUAAUCUGC --..............-..................((((((((((((.......))))).........((......)).................)))))))... ( -15.00, z-score = -1.57, R) >droSec1.super_20 227578 103 - 1148123 --UAACCCUAUAUAGUCCAAAAUACCCAUACUACCGAUUGUCACGAGAAUCAUACUCGUAAUACCAUCGCGCCCCAGCAAUCACUAAUCGUAAAAGAUAAUCUGC --.................................((((((((((((.......))))).........((......)).................)))))))... ( -15.00, z-score = -2.53, R) >droYak2.chrX 7872435 94 - 21770863 UAUAACCCUCUACUGC-CG----CCUAAUACUACCGAUUGUCACGUGA------UGCGUAAUACUAACGCGCCGCAGCAAUCACUAAUCGUAAACGAUAAUCUGU ................-..----............((((((...(((.------(((((.......))))).))).))))))....((((....))))....... ( -17.10, z-score = -1.21, R) >droEre2.scaffold_4690 5453602 89 + 18748788 --UAACCCUAUAUAGC--------CUAAUACUACCGAAUGUCACGUGAA------UCGUAAUACAAUCGCGCCUCAGCCGUCACUAAUCGUAAAAGAUAAUCUCU --..............--------..........(((..((.((((((.------.(((.........)))..)))..))).))...)))............... ( -6.70, z-score = 1.04, R) >consensus __UAACCCUAUAUAGU_CGAAAUACCCAUACUACCGAUUGUCACGAGAAUCAUACUCGUAAUACCAUCGCGCCCCAGCAAUCACUAAUCGUAAAAGAUAAUCUGC ...................................((((((((((((.......))))).........((......)).................)))))))... (-10.80 = -11.44 + 0.64)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:40:53 2011