
| Sequence ID | dm3.chrX |
|---|---|
| Location | 13,285,315 – 13,285,425 |
| Length | 110 |
| Max. P | 0.908848 |

| Location | 13,285,315 – 13,285,425 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 135 |
| Reading direction | forward |
| Mean pairwise identity | 50.50 |
| Shannon entropy | 0.85866 |
| G+C content | 0.42569 |
| Mean single sequence MFE | -30.18 |
| Consensus MFE | -8.06 |
| Energy contribution | -7.67 |
| Covariance contribution | -0.38 |
| Combinations/Pair | 1.60 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.27 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.20 |
| SVM RNA-class probability | 0.908848 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
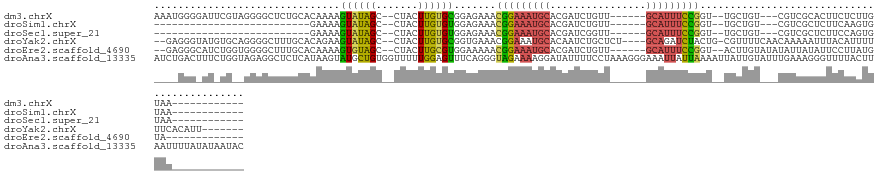
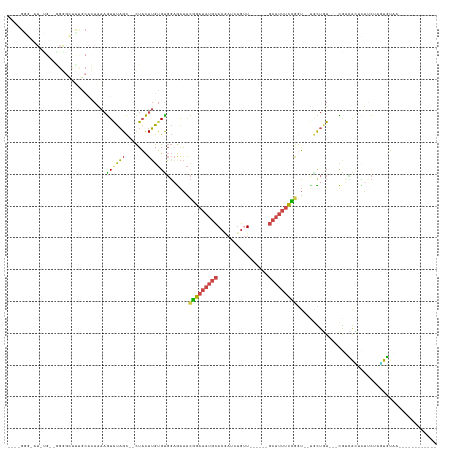
>dm3.chrX 13285315 110 + 22422827 AAAUGGGGAUUCGUAGGGGCUCUGCACAAAAGUAUAGC--CUACUUGUGCGGAGAAACGGAAAUGCACGAUCUGUU------GCAUUUCCGGU--UGCUGU---CGUCGCACUUCUCUUGUAA------------ ....(((((.......(((((.(((......))).)))--))....(((((..((..((((((((((........)------)))))))))..--.....)---)..))))).))))).....------------ ( -37.80, z-score = -2.60, R) >droSim1.chrX 10177753 84 + 17042790 --------------------------GAAAAGUAUAGC--CUACUUGUGUGGAGAAACGGAAAUGCACGAUCUGUU------GCAUUUCCGGU--UGCUGU---CGUCGCUCUUCAAGUGUAA------------ --------------------------(((.((((((((--((((....)))).....((((((((((........)------)))))))))..--.)))))---....))).)))........------------ ( -25.50, z-score = -2.33, R) >droSec1.super_21 1038753 84 + 1102487 --------------------------GAAAAGUAUAGC--CUACUUGUGUGGAGAAACGGAAAUGCACGAUCGGUU------GCAUUUCCGGU--UGCUGU---CGUCGCUCUUCCAGUGUAA------------ --------------------------(((.((((((((--((((....)))).....((((((((((........)------)))))))))..--.)))))---....))).)))........------------ ( -24.60, z-score = -1.51, R) >droYak2.chrX 7565637 119 + 21770863 --GAGGGUAUGUGCAGGGGCUUUGCACAGAAGUAUAGC--CUACUUGUGCGGUGAAACGGAAAUGCACAAUCUGCUCU----GCAGAUCUACUG-CGUUUUCAACAAAAAUUUACAUUUUUUCACAUU------- --.......((..((((((((.(((......))).)))--)..))))..))(((((..(((((((((..((((((...----))))))....))-)))))))....(((((....))))))))))...------- ( -35.10, z-score = -2.26, R) >droEre2.scaffold_4690 5165372 110 - 18748788 --GAGGGCAUCUGGUGGGGCUUUGCACAAAAGUGUAGC--CUACUUGCGUGGAAAAACGGAAAUGCACGAUCUGUU------GCAUUUCCGGU--ACUUGUAUAUAUUAUAUUCCUUAUGUA------------- --(((((((...((((((.((..((((....)))))))--))))))))(((.((...((((((((((........)------)))))))))..--..)).)))..........)))).....------------- ( -33.10, z-score = -1.91, R) >droAna3.scaffold_13335 2484898 135 - 3335858 AUCUGACUUUCUGGUAGAGGCUCUCAUAAGUAUGCUGUGGUUUUUGGAGUUUCAGGGUAGAAAAGGAUAUUUUCCUAAAGGGAAAUUAUUAAAAUUAUUGUAUUUGAAAGGGUUUUACUUAAUUUUAUAUAAUAC ((((...((((((.(.((((((((...((.(((...))).))...))))))))..).)))))).))))(((((((....))))))).........((((((((..((..(((.....)))..))..)))))))). ( -25.00, z-score = 0.11, R) >consensus ____GGG_AU_UG__GGGGCUUUGCACAAAAGUAUAGC__CUACUUGUGUGGAGAAACGGAAAUGCACGAUCUGUU______GCAUUUCCGGU__UGCUGU___CGUCACUCUUCUAGUGUAA____________ ...............................((((((.......)))))).......(((((((((................)))))))))............................................ ( -8.06 = -7.67 + -0.38)
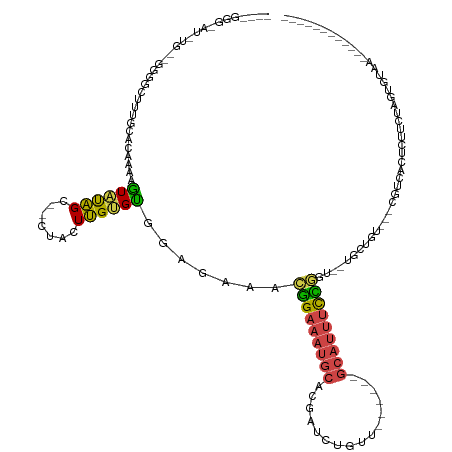

| Location | 13,285,315 – 13,285,425 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 135 |
| Reading direction | reverse |
| Mean pairwise identity | 50.50 |
| Shannon entropy | 0.85866 |
| G+C content | 0.42569 |
| Mean single sequence MFE | -22.92 |
| Consensus MFE | -5.61 |
| Energy contribution | -6.36 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.24 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.95 |
| SVM RNA-class probability | 0.859624 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
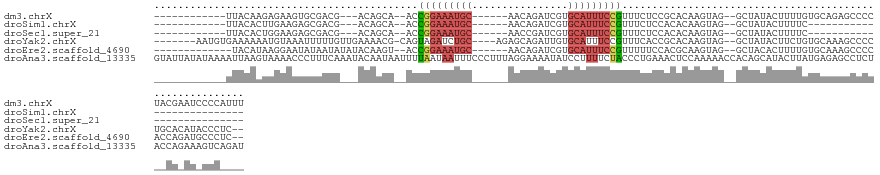
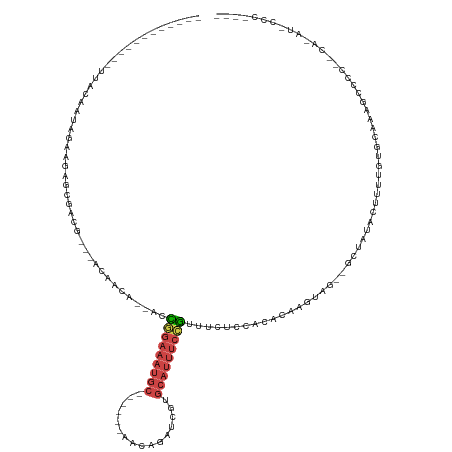
>dm3.chrX 13285315 110 - 22422827 ------------UUACAAGAGAAGUGCGACG---ACAGCA--ACCGGAAAUGC------AACAGAUCGUGCAUUUCCGUUUCUCCGCACAAGUAG--GCUAUACUUUUGUGCAGAGCCCCUACGAAUCCCCAUUU ------------......(((((((((....---...)))--..(((((((((------(........)))))))))))))))).(((((((.((--......)))))))))....................... ( -31.50, z-score = -3.32, R) >droSim1.chrX 10177753 84 - 17042790 ------------UUACACUUGAAGAGCGACG---ACAGCA--ACCGGAAAUGC------AACAGAUCGUGCAUUUCCGUUUCUCCACACAAGUAG--GCUAUACUUUUC-------------------------- ------------....(((((....((....---...)).--..(((((((((------(........))))))))))..........)))))..--............-------------------------- ( -20.60, z-score = -2.36, R) >droSec1.super_21 1038753 84 - 1102487 ------------UUACACUGGAAGAGCGACG---ACAGCA--ACCGGAAAUGC------AACCGAUCGUGCAUUUCCGUUUCUCCACACAAGUAG--GCUAUACUUUUC-------------------------- ------------......((((.((((....---...)).--..(((((((((------(........))))))))))..))))))...(((((.--....)))))...-------------------------- ( -22.20, z-score = -2.46, R) >droYak2.chrX 7565637 119 - 21770863 -------AAUGUGAAAAAAUGUAAAUUUUUGUUGAAAACG-CAGUAGAUCUGC----AGAGCAGAUUGUGCAUUUCCGUUUCACCGCACAAGUAG--GCUAUACUUCUGUGCAAAGCCCCUGCACAUACCCUC-- -------...(((((((((((((....(((((((....)(-(((.....))))----..))))))...)))))))...)))))).(((((((((.--....))))..)))))...((....))..........-- ( -26.70, z-score = -0.85, R) >droEre2.scaffold_4690 5165372 110 + 18748788 -------------UACAUAAGGAAUAUAAUAUAUACAAGU--ACCGGAAAUGC------AACAGAUCGUGCAUUUCCGUUUUUCCACGCAAGUAG--GCUACACUUUUGUGCAAAGCCCCACCAGAUGCCCUC-- -------------..(((..((..................--..(((((((((------(........)))))))))).......((....)).(--(((.(((....)))...))))...))..))).....-- ( -26.60, z-score = -2.62, R) >droAna3.scaffold_13335 2484898 135 + 3335858 GUAUUAUAUAAAAUUAAGUAAAACCCUUUCAAAUACAAUAAUUUUAAUAAUUUCCCUUUAGGAAAAUAUCCUUUUCUACCCUGAAACUCCAAAAACCACAGCAUACUUAUGAGAGCCUCUACCAGAAAGUCAGAU .................(((.....((((((...................(((((.....))))).......((((......)))).......................))))))....)))............. ( -9.90, z-score = 0.68, R) >consensus ____________UUACAAUAGAAGAGCGACG___ACAACA__ACCGGAAAUGC______AACAGAUCGUGCAUUUCCGUUUCUCCACACAAGUAG__GCUAUACUUUUGUGCAAAGCCCC__CA_AU_CCC____ ............................................(((((((((................)))))))))......................................................... ( -5.61 = -6.36 + 0.75)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:40:00 2011