
| Sequence ID | dm3.chrX |
|---|---|
| Location | 13,257,059 – 13,257,118 |
| Length | 59 |
| Max. P | 0.946430 |

| Location | 13,257,059 – 13,257,118 |
|---|---|
| Length | 59 |
| Sequences | 9 |
| Columns | 71 |
| Reading direction | reverse |
| Mean pairwise identity | 74.53 |
| Shannon entropy | 0.48990 |
| G+C content | 0.32271 |
| Mean single sequence MFE | -12.17 |
| Consensus MFE | -5.34 |
| Energy contribution | -5.38 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.64 |
| Mean z-score | -2.35 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.52 |
| SVM RNA-class probability | 0.946430 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
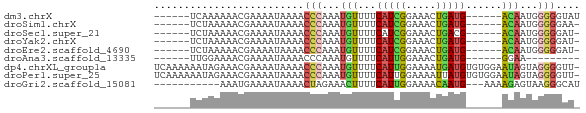
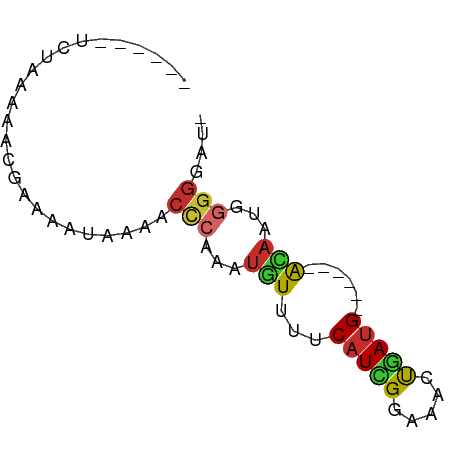
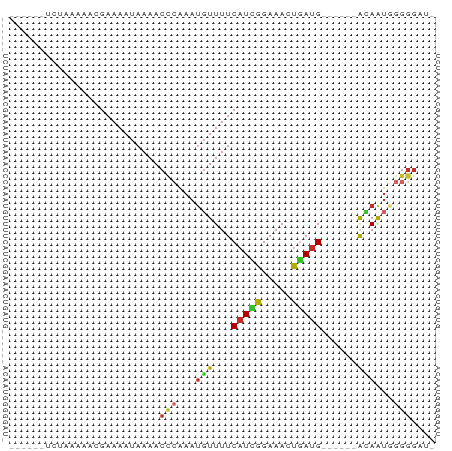
>dm3.chrX 13257059 59 - 22422827 ------UCAAAAAACGAAAAUAAAACCCAAAUGUUUUCAUCGGAAACUGAUG------ACAAUGGGGGUAU ------..................((((...((((.((((((....).))))------).)))).)))).. ( -14.80, z-score = -3.06, R) >droSim1.chrX 10148267 58 - 17042790 ------UCUAAAAACGAAAAUAAAACCCAAAUGUUUUCAUCGGAAACUGAUG------ACAAUGGGGGAA- ------...................(((...((((.((((((....).))))------).)))).)))..- ( -13.70, z-score = -2.73, R) >droSec1.super_21 1010730 58 - 1102487 ------UCUAAAAACGAAAAUAAAACCCAAAUGUUUUCAUCGGAAACUGACG------ACAAUGGGGGAU- ------...................(((...((((.(((..(....)))).)------)))...)))...- ( -11.70, z-score = -1.79, R) >droYak2.chrX 7537797 58 - 21770863 ------UCUAAAAACGAAAAUAAAACCCAAAUGUUUUCAUCGGAAACUGAUG------ACAAUGGGGGAU- ------...................(((...((((.((((((....).))))------).)))).)))..- ( -13.70, z-score = -2.75, R) >droEre2.scaffold_4690 5135916 58 + 18748788 ------UCUAAAAACGAAAAUAAAACCCAAAUGUUUUCAUCGGAAACUGAUG------ACAAUGGGGGAU- ------...................(((...((((.((((((....).))))------).)))).)))..- ( -13.70, z-score = -2.75, R) >droAna3.scaffold_13335 2449408 50 + 3335858 ------UUGGAAAACGAAAAUAAAACCCAAAUGUUUUCAUUGGAAACUGAUG------GGAA--------- ------...................((((...(((((.....)))))...))------))..--------- ( -9.70, z-score = -1.63, R) >dp4.chrXL_group1a 3152850 70 + 9151740 UCAAAAAAUAGAAACGAAAAUAAAACCCAAAUGUUUUCAUUGGAAAAUGAUGUGUGGAAUAGUAGGGGUU- .......................(((((...((((((((((....)))))......)))))....)))))- ( -11.30, z-score = -1.38, R) >droPer1.super_25 1375225 70 - 1448063 UCAAAAAAUAGAAACGAAAAUAAAACCCAAAUGUUUUCAUUGGAAAAUUAUGUGUGGAAUAGUAGGGGUU- .......................(((((......((((((.............))))))......)))))- ( -9.12, z-score = -0.30, R) >droGri2.scaffold_15081 3057490 57 + 4274704 -----------AAAUGAAAAUAAAACUAGAAACUUUUCAUUGGAAAACAAUG---AAAAGAGUAAGGGCAU -----------.....................((((((((((.....)))))---)))))........... ( -11.80, z-score = -4.77, R) >consensus ______UCUAAAAACGAAAAUAAAACCCAAAUGUUUUCAUCGGAAACUGAUG______ACAAUGGGGGAU_ .........................(((...(((...(((((.....)))))......)))...))).... ( -5.34 = -5.38 + 0.04)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:39:56 2011