
| Sequence ID | dm3.chrX |
|---|---|
| Location | 13,247,650 – 13,247,737 |
| Length | 87 |
| Max. P | 0.655086 |

| Location | 13,247,650 – 13,247,737 |
|---|---|
| Length | 87 |
| Sequences | 6 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 85.42 |
| Shannon entropy | 0.27439 |
| G+C content | 0.55241 |
| Mean single sequence MFE | -32.08 |
| Consensus MFE | -21.48 |
| Energy contribution | -22.90 |
| Covariance contribution | 1.42 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.655086 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
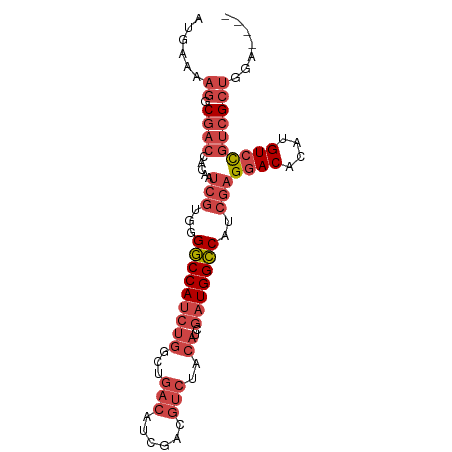
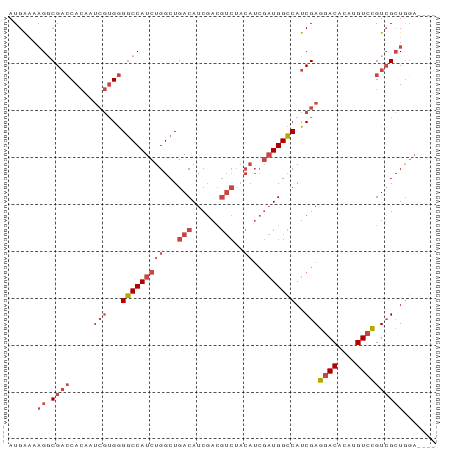
>dm3.chrX 13247650 87 + 22422827 AUGAAAAGGCGACCACAAUCGUGGGGCCAUCUGGCUGACAUCGACGUCUACAUCGAUGGCCAUCGAGGACACAUGUCCGUCGCUGGA---- .......((((((.....(((...(((((((..(..(((......)))..)...)))))))..)))((((....))))))))))...---- ( -35.20, z-score = -2.43, R) >droSim1.chrX 10139936 87 + 17042790 AUGAAAAGGCGACCACAAUCGUGGGGCCAUCUGGCUGACAUCGACGUCUACAUCGAUGGCCAUCGAGGACACAUGUCCGUCGCUGGA---- .......((((((.....(((...(((((((..(..(((......)))..)...)))))))..)))((((....))))))))))...---- ( -35.20, z-score = -2.43, R) >droSec1.super_21 1002465 87 + 1102487 AUGAAAAGGCGACCACAAUCGUGGGGCCAUCUGGCUGACAUCGACGUCUACAUCGAUGGCCAUCGAGGACACAUGUCCGUCGCUGGA---- .......((((((.....(((...(((((((..(..(((......)))..)...)))))))..)))((((....))))))))))...---- ( -35.20, z-score = -2.43, R) >droYak2.chrX 7527406 90 + 21770863 AUGAAAAG-CGACCACAAUCGUGGGGCCAUCUGGCUGACAUCGACGUCUACAUCGAUGGCCAUCGAGGACACAUGUCCGUCGCUGCCUGGA ......((-((((.....(((...(((((((..(..(((......)))..)...)))))))..)))((((....))))))))))....... ( -35.20, z-score = -2.15, R) >droEre2.scaffold_4690 5122498 86 - 18748788 AUGAAAUG-CGACCACAAUCGUGGGGCCAUCUGGCUGACAUCGACGUCUACAUCGAUGGCCAUCGAGGACACAUGUCCGUCGCUGGA---- .......(-((((.....(((...(((((((..(..(((......)))..)...)))))))..)))((((....)))))))))....---- ( -33.80, z-score = -2.07, R) >droPer1.super_25 1365854 70 + 1448063 AUGGGAAGGCG--CAGAAA---GAGUCCAUCUGGCUGACAUCUA------------UGGGCAUCGAGUACACAUGUCUGUGGAUAGA---- ...(...(((.--((((..---.......)))))))..)(((((------------(((((((.(......)))))))))))))...---- ( -17.90, z-score = 0.14, R) >consensus AUGAAAAGGCGACCACAAUCGUGGGGCCAUCUGGCUGACAUCGACGUCUACAUCGAUGGCCAUCGAGGACACAUGUCCGUCGCUGGA____ .........(((((((....))))(((((((..(..(((......)))..)...))))))).))).((((....))))............. (-21.48 = -22.90 + 1.42)

| Location | 13,247,650 – 13,247,737 |
|---|---|
| Length | 87 |
| Sequences | 6 |
| Columns | 91 |
| Reading direction | reverse |
| Mean pairwise identity | 85.42 |
| Shannon entropy | 0.27439 |
| G+C content | 0.55241 |
| Mean single sequence MFE | -30.65 |
| Consensus MFE | -20.90 |
| Energy contribution | -23.48 |
| Covariance contribution | 2.58 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.565133 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
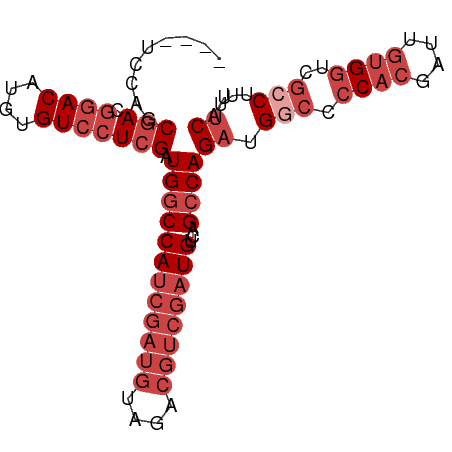
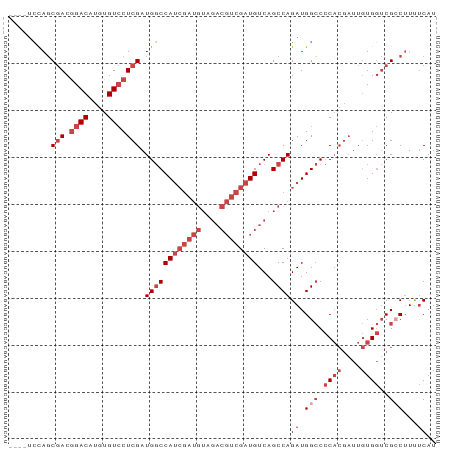
>dm3.chrX 13247650 87 - 22422827 ----UCCAGCGACGGACAUGUGUCCUCGAUGGCCAUCGAUGUAGACGUCGAUGUCAGCCAGAUGGCCCCACGAUUGUGGUCGCCUUUUCAU ----.....(((.((((....))))))).((((((((((((....))))))))...))))((.(((.((((....))))..)))...)).. ( -34.40, z-score = -2.38, R) >droSim1.chrX 10139936 87 - 17042790 ----UCCAGCGACGGACAUGUGUCCUCGAUGGCCAUCGAUGUAGACGUCGAUGUCAGCCAGAUGGCCCCACGAUUGUGGUCGCCUUUUCAU ----.....(((.((((....))))))).((((((((((((....))))))))...))))((.(((.((((....))))..)))...)).. ( -34.40, z-score = -2.38, R) >droSec1.super_21 1002465 87 - 1102487 ----UCCAGCGACGGACAUGUGUCCUCGAUGGCCAUCGAUGUAGACGUCGAUGUCAGCCAGAUGGCCCCACGAUUGUGGUCGCCUUUUCAU ----.....(((.((((....))))))).((((((((((((....))))))))...))))((.(((.((((....))))..)))...)).. ( -34.40, z-score = -2.38, R) >droYak2.chrX 7527406 90 - 21770863 UCCAGGCAGCGACGGACAUGUGUCCUCGAUGGCCAUCGAUGUAGACGUCGAUGUCAGCCAGAUGGCCCCACGAUUGUGGUCG-CUUUUCAU ....(((..(((.((((....))))))).((((.(((((((....)))))))))))))).((.(((.((((....))))..)-))..)).. ( -35.10, z-score = -1.85, R) >droEre2.scaffold_4690 5122498 86 + 18748788 ----UCCAGCGACGGACAUGUGUCCUCGAUGGCCAUCGAUGUAGACGUCGAUGUCAGCCAGAUGGCCCCACGAUUGUGGUCG-CAUUUCAU ----....(((((((((....))))(((.(((.((((((((....))))))))...(((....))).)))))).....))))-)....... ( -34.20, z-score = -2.22, R) >droPer1.super_25 1365854 70 - 1448063 ----UCUAUCCACAGACAUGUGUACUCGAUGCCCA------------UAGAUGUCAGCCAGAUGGACUC---UUUCUG--CGCCUUCCCAU ----..........(((((.(((...........)------------)).))))).((((((.......---..))))--.))........ ( -11.40, z-score = 0.61, R) >consensus ____UCCAGCGACGGACAUGUGUCCUCGAUGGCCAUCGAUGUAGACGUCGAUGUCAGCCAGAUGGCCCCACGAUUGUGGUCGCCUUUUCAU .........(((.((((....))))))).((((((((((((....))))))))...))))((.(((.((((....))))..)))...)).. (-20.90 = -23.48 + 2.58)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:39:54 2011