
| Sequence ID | dm3.chrX |
|---|---|
| Location | 13,195,517 – 13,195,645 |
| Length | 128 |
| Max. P | 0.526843 |

| Location | 13,195,517 – 13,195,607 |
|---|---|
| Length | 90 |
| Sequences | 3 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 65.47 |
| Shannon entropy | 0.48658 |
| G+C content | 0.30263 |
| Mean single sequence MFE | -16.66 |
| Consensus MFE | -8.41 |
| Energy contribution | -8.63 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.526843 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 13195517 90 - 22422827 AUCUCAAAGUAUUGCCUACUUGUUUAGGCAAUAAAAGUAUUUCCGAGAAUUACAUACGUUCUUUAUGGAUACAAGGUAUUUAUUAGAAUA---- .((((....(((((((((......)))))))))...(......))))).........(((((..((((((((...)))))))).))))).---- ( -20.60, z-score = -2.87, R) >droEre2.scaffold_4690 5071060 92 + 18748788 AUCUCAAAGUGUUGCCUACUACUUAAGGUAAUAAAAGUAUUUCCAACAAACAAGAUCUUA-AUGG-GGGUAUAUGGUAUUUAUGUAUUUGCAAG ....((((.((((((((........))))))))...((((..(((...............-.)))-..))))..............)))).... ( -14.79, z-score = 0.22, R) >droYak2.chrX 7472919 79 - 21770863 AUCUCAAAUUGUUGCCUACUUGUUUAGGCUAUAAAAGUAUUUCCGAGAAAUACGAUCAUACAUGAUAUUUAUUUGGUAG--------------- ...(((((((((.(((((......))))).))))..(((((((...))))))).((((....)))).....)))))...--------------- ( -14.60, z-score = -1.43, R) >consensus AUCUCAAAGUGUUGCCUACUUGUUUAGGCAAUAAAAGUAUUUCCGAGAAAUACGAUCAUACAUGAUGGAUAUAUGGUAUUUAU_____U_____ .........(((((((((......)))))))))............................................................. ( -8.41 = -8.63 + 0.23)



| Location | 13,195,541 – 13,195,645 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 61.80 |
| Shannon entropy | 0.67669 |
| G+C content | 0.31737 |
| Mean single sequence MFE | -15.82 |
| Consensus MFE | -5.32 |
| Energy contribution | -6.38 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.33 |
| Structure conservation index | 0.34 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.521208 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 13195541 104 - 22422827 CAACUAAGUUAACAGACUACAAUCUACGGAUAUUAGCGAUCUCAAAGUAUUGCCUACUUGUUUAGGCAAUAAAAGUAUUUCCGAGAAUUACAUACGUUCUUUAU ......((((....))))...(((....)))...((((.........(((((((((......)))))))))...(((.(((...))).)))...))))...... ( -18.60, z-score = -1.40, R) >droEre2.scaffold_4690 5071090 100 + 18748788 GAAAUAAAUUAACACACUAUAAUCUACGGAUAUUAGCGAUCUCAAAGUGUUGCCUACUACUUAAGGUAAUAAAAGUAUUUCCAACAAACAAGAUCUUAAU---- ((((((........(.(((..(((....)))..))).).........((((((((........))))))))....))))))...................---- ( -15.80, z-score = -2.23, R) >droYak2.chrX 7472941 85 - 21770863 --------GAAACAGA-AAUAAAC-AGAAAUAUUAGCGAUCUCAAAUUGUUGCCUACUUGUUUAGGCUAUAAAAGUAUUUCCGAGAAAUACGAUC--------- --------........-.......-............((((.....((((.(((((......))))).))))..(((((((...)))))))))))--------- ( -15.80, z-score = -2.09, R) >droSec1.super_21 951097 94 - 1102487 CAACUAAGUUAACAGACUACAAUCUACGGAUAUUAGCGAUCUCAAAGUAUUGCCUACUUGUUUAGGCAGUAAAAGUAUUUCCGAGAAAUACGUU---------- ..............(.(((..(((....)))..))))..........(((((((((......)))))))))...(((((((...)))))))...---------- ( -19.80, z-score = -1.77, R) >droWil1.scaffold_180702 4041110 83 + 4511350 ----UAUGUUACUUAG--AAAAACUAUUUAGUUAGGACAUCCCUAAAUAUC-----CUUUUGUAG---ACAAAGAUCUUUUGUACAGAUUUCCGUGU------- ----..(((.((..((--(.....(((((((...((....)))))))))..-----.(((((...---.))))).)))...)))))...........------- ( -9.10, z-score = 0.83, R) >consensus _AA_UAAGUUAACAGACUACAAUCUACGGAUAUUAGCGAUCUCAAAGUAUUGCCUACUUGUUUAGGCAAUAAAAGUAUUUCCGAGAAAUACGAUC_U_______ ...............................................(((((((((......)))))))))................................. ( -5.32 = -6.38 + 1.06)

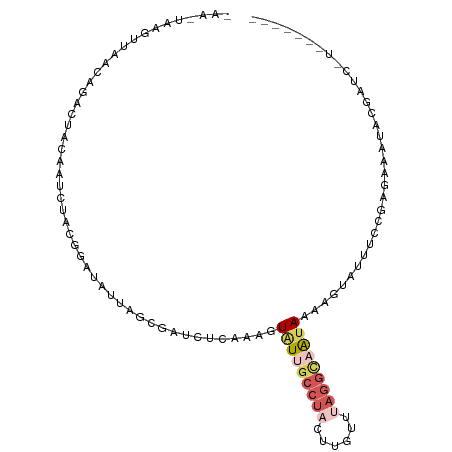

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:39:44 2011