| Sequence ID | dm3.chrX |
|---|---|
| Location | 13,027,927 – 13,028,026 |
| Length | 99 |
| Max. P | 0.880016 |

| Location | 13,027,927 – 13,028,026 |
|---|---|
| Length | 99 |
| Sequences | 15 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 72.69 |
| Shannon entropy | 0.57204 |
| G+C content | 0.56680 |
| Mean single sequence MFE | -35.31 |
| Consensus MFE | -12.14 |
| Energy contribution | -12.04 |
| Covariance contribution | -0.10 |
| Combinations/Pair | 1.71 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.34 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.04 |
| SVM RNA-class probability | 0.880016 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 13027927 99 + 22422827 -----------------ACUUUGACAUCCAGUCGAAUGCUCUGCUCAAUGCGGUGCCGUAUCUAACCUCCUGGUUCGUGGGCAUUGCCUGCUCCGCCCUGGCGGAUUGGAUGCUAG- -----------------..((((((.....)))))).((.((((.....)))).)).((((((((((....)))..(..(((...)))..)(((((....))))).)))))))...- ( -36.90, z-score = -2.57, R) >droSim1.chrX 9917525 99 + 17042790 -----------------ACUUUGACAUCCAGUCGAAUGCUCUGCUCAAUGCGGUGCCGUAUCUAACCUCCUGGUUCGUGGGCAUUGCCUGCUCCGCCCUGGCGGAUUGGAUGCUAG- -----------------..((((((.....)))))).((.((((.....)))).)).((((((((((....)))..(..(((...)))..)(((((....))))).)))))))...- ( -36.90, z-score = -2.57, R) >droSec1.super_21 780040 99 + 1102487 -----------------ACUUUGACAUCCAGUCGAAUGCUCUGCUCAAUGCGGUGCCGUAUCUAACCUCCUGGUUCGUGGGCAUUGCCUGUUCCGCCCUGGCGGAUUGGAUGCUAG- -----------------.......(((((((((((((((..(((((((((((....)))))..((((....))))..))))))..))..))))(((....))))))))))))....- ( -37.00, z-score = -2.79, R) >droYak2.chrX 7303207 99 + 21770863 -----------------ACUUUGACAUCCAGUCGAAUGCUCUGCUGAAUGCGGUGCCGUAUCUAACCGCCUGGUUCGUGGGCAUCGCCUGCUCCGCUCUGGCGGAUUGGAUGCUGG- -----------------..((((((.....)))))).((.((((.....)))).)).(((((((((((((.((..((.(((((.....)))))))..))))))).))))))))...- ( -40.50, z-score = -2.84, R) >droEre2.scaffold_4690 4903453 99 - 18748788 -----------------ACUUUGACAUCCAGUCCAAUGCUCUGCUGAACGCGGUGCCGUAUUUAACCGCCUGGUUCGUGGGCAUUGCCUGCUCCGCCCUGGCGGAUUGGAUGCUGG- -----------------.......(((((((((((((((.(..(.(((((((((..........)))))...)))))..))))))(((.((...))...)))))))))))))....- ( -40.10, z-score = -2.67, R) >droAna3.scaffold_13117 2350191 99 + 5790199 -----------------ACUUUGACAUCCAGUCGAACGCCCUGCUGAACGCCGUUCCCUACCUGACCGCCUGGUUCGUGGGCAUAGCCUGCUCCGCCCUCGCCGACUGGAUGCUCG- -----------------.....((((((((((((((((....((.....))))))).......((((....))))((.((((..((....))..)))).))..))))))))).)).- ( -33.90, z-score = -2.80, R) >dp4.chrXL_group1e 5374237 99 + 12523060 -----------------ACUUUGACAUCCAGUCGAAUGCCCUGCUCAACGCCGUGCCCUACCUCACGGCCUGGUUUGUGGGUAUCGCCUGCUCUGCCCUGGCCGACUGGAUGCUGG- -----------------.......((((((((((...(((.........((((((........))))))..(((..(..((.....))..)...)))..)))))))))))))....- ( -39.40, z-score = -3.39, R) >droPer1.super_21 286132 99 + 1645244 -----------------ACUUUGACAUCCAGUCGAAUGCCCUGCUCAACGCCGUGCCCUACCUCACGGCCUGGUUUGUGGGUAUCGCCUGCUCUGCCCUGGCCGACUGGAUGCUGG- -----------------.......((((((((((...(((.........((((((........))))))..(((..(..((.....))..)...)))..)))))))))))))....- ( -39.40, z-score = -3.39, R) >droWil1.scaffold_181150 1793866 99 + 4952429 -----------------AUUUCGACAUCCAAUCGAAUGCCAUCCUAAACGCUGUGCCCCCGCUAACGGCCUGGUUUGUGGGCAUUGCCUGUUCCGCGCUCGCUGAUUGGAUGCUGG- -----------------....((.((((((((((((((((....((((((((((((....))..)))))...)))))..)))))).........((....)))))))))))).)).- ( -30.70, z-score = -0.90, R) >droVir3.scaffold_12928 553001 99 + 7717345 -----------------ACUUUGACAUCCAAUCGAAUGCGCUGCUCAAUGCGGUGCCGUUUCUAACCUCCUGGUUCAUGGGCAUCGCCUGCUCCGCCCUGGCGGAUUGGAUGCUCG- -----------------.....((((((((((((((.(((((((.....)))))))...))).((((....))))...(((((.....)))))(((....)))))))))))).)).- ( -38.80, z-score = -3.32, R) >droMoj3.scaffold_6328 1513266 99 + 4453435 -----------------ACUUUGACAUCCAGUCGAAUGCCCUGCUCAAUGCCGUGCCGUAUCUGACCGCCUGGUUCGUGGGCAUCGCCUGCUCCGCCCUGGCGGACUGGAUGCUCG- -----------------.....(((((((((((....((..((((((..((((.((.((.....)).)).))))...))))))..))......(((....)))))))))))).)).- ( -39.90, z-score = -2.91, R) >droGri2.scaffold_14853 2237168 99 - 10151454 -----------------ACUUUGACAUCCAGUCGAAUGCACUGCUGAAUGCCGUACCCUAUCUGACCUCCUGGUUCAUGGGCAUCGCGUGCUCCGCUCUGGCGGACUGGAUGCUCG- -----------------.....(((((((((((....((((.((.((.(((((........).((((....))))....))))))))))))..(((....)))))))))))).)).- ( -36.40, z-score = -2.91, R) >anoGam1.chr3L 38648147 117 + 41284009 UAACGUUGACAUCUUCUUUUCCCCCAUUUUCCAGAACGGUGUCCUUUCGGCCCUGCCCUACCUGGCGAUGUGGCUGUUCUCGAUCGUGGUCGGCUGGGUGGCCGACUGGAUGCUGAC ....................................((((((((..((((((..((((...(((((.(((...(.......)..))).)))))..))))))))))..)))))))).. ( -38.80, z-score = -0.96, R) >apiMel3.GroupUn 39575411 99 + 399230636 -----------------AUUUCGAUGUACAACAGGAUGCAUUCUUUUCAGCACUGCCUUAUUUAAGUUCUUGGUUAGUCGGUCUCGGUAUUUCAAGUUUCGCAGAUGCUCUCCUUG- -----------------.....((((((........))))))......(((((((((((....)))..(((((....(((....)))....)))))....)))).)))).......- ( -15.00, z-score = 1.09, R) >triCas2.ChLG8 12669400 99 + 15773733 -----------------ACUUUGACAUUCAAGCUAACGCAUUUUUGUCCGCAAUUCCGUAUCUUACAAGUUGGUGGUGCGGGAUUCUUAUCAGUUUCGUGGCCGAUUGGCUGUUGA- -----------------...(..(((..(((((....))....(((.((((....(((((((...(.....)..)))))))((..((....))..)))))).))))))..)))..)- ( -26.00, z-score = -0.83, R) >consensus _________________ACUUUGACAUCCAGUCGAAUGCUCUGCUCAAUGCCGUGCCGUAUCUAACCGCCUGGUUCGUGGGCAUCGCCUGCUCCGCCCUGGCGGAUUGGAUGCUGG_ ........................(((((((......((...((.....((.(((((......(((......)))....))))).)).......))....))...)))))))..... (-12.14 = -12.04 + -0.10)


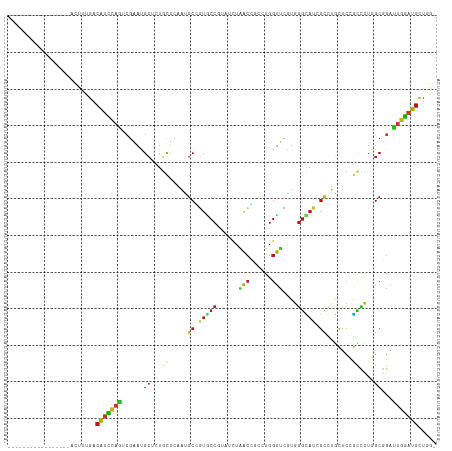
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:39:17 2011