
| Sequence ID | dm3.chrX |
|---|---|
| Location | 12,956,085 – 12,956,190 |
| Length | 105 |
| Max. P | 0.999407 |

| Location | 12,956,085 – 12,956,190 |
|---|---|
| Length | 105 |
| Sequences | 11 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 70.86 |
| Shannon entropy | 0.56731 |
| G+C content | 0.46641 |
| Mean single sequence MFE | -32.91 |
| Consensus MFE | -22.24 |
| Energy contribution | -21.32 |
| Covariance contribution | -0.93 |
| Combinations/Pair | 1.38 |
| Mean z-score | -2.65 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.86 |
| SVM RNA-class probability | 0.999407 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
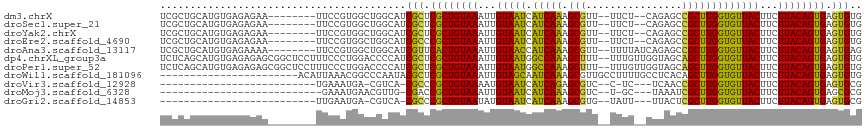
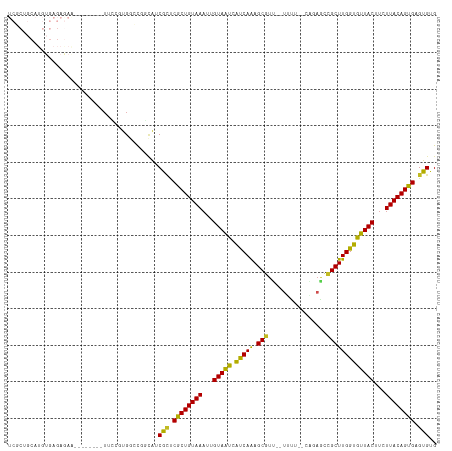
>dm3.chrX 12956085 105 + 22422827 UCGCUGCAUGUGAGAGAA--------UUCCGUGGCUGGCAUCGCUCGCUGUAAAUUGUAAUCAUCAAAGCGUU--UUCU--CAGAGCCGCUUGGUGUUACUUCUUACAGUGAGUGUG ..((((((((.(((....--------)))))).)).)))..((((((((((((...((((.((((((.(((((--(...--..))).)))))))))))))...)))))))))))).. ( -36.40, z-score = -2.66, R) >droSec1.super_21 701213 105 + 1102487 UCGCUGCAUGUGAGAGAA--------UUCCGUGGCUGGCAUCGCUCGCUGUAAAUUGUAAUCAUCAAAGCGUU--UUCU--CAGAGCCGCUUGGUGUUACUUCUUACAGUGAGUGUG ..((((((((.(((....--------)))))).)).)))..((((((((((((...((((.((((((.(((((--(...--..))).)))))))))))))...)))))))))))).. ( -36.40, z-score = -2.66, R) >droYak2.chrX 7229765 105 + 21770863 UCGCUGCAUGUGAGAGAA--------UUCCGUGGCUGGCAUCGCUCGCUGUAAAUUGUAAUCAUCAAAGCGUU--UUCU--CAGAGCCGCUUGGUGUUACUUCUUACAGUGAGUGUG ..((((((((.(((....--------)))))).)).)))..((((((((((((...((((.((((((.(((((--(...--..))).)))))))))))))...)))))))))))).. ( -36.40, z-score = -2.66, R) >droEre2.scaffold_4690 4833925 105 - 18748788 UCGCUGCAUGUGAGAGAA--------UUCCGUGGCUGGCAUCGCCCGCUGUAAAUUGUAAUCAUCAAAGCGUU--UUCU--CAGAGCCGCUUGGUGUUACUUCUUACAGUGAGUGUG ..((((((((.(((....--------)))))).)).)))..(((.((((((((...((((.((((((.(((((--(...--..))).)))))))))))))...)))))))).))).. ( -31.60, z-score = -1.25, R) >droAna3.scaffold_13117 3891779 107 + 5790199 UCGCUGCAUGUGAGAAAA--------UUCCGUGGCUGGCAUCGUUCACUGUAAAUUGUAACCAUCAAAGCGUU--UUUUAUCAGAGCCGCUUGGUGUUACUUCUUACAGUGAGUGAG ..((((((((.(((....--------)))))).)).))).(((((((((((((...(((((.(((((.(((.(--((......))).)))))))))))))...))))))))))))). ( -34.20, z-score = -2.78, R) >dp4.chrXL_group3a 1299084 115 - 2690836 UCUCAGCAUGUGAGAGAGCGGCUCCUUUCCCUGGACCCCAUCGCUCGCUGUAAAUUGUAAUGGCCAAAGCUUU--UUUGUUGGUAGCAGCUUGGUGUUACUUCUUACAGUGAGUGUG (((((.....)))))..(.((.(((.......))).)))..((((((((((((...((((((.((((.(((.(--(........)).)))))))))))))...)))))))))))).. ( -37.30, z-score = -1.84, R) >droPer1.super_52 413302 115 + 549863 UCUCAGCAUGUGAGAGAGCGGCUCCUUUCCCUGGACCCCAUCGCUCGCUGUAAAUUGUAAUGGCCAAAGCUUU--UUUGUUGGUAGCAGCUUGGUGUUACUUCUUACAGUGAGUGUG (((((.....)))))..(.((.(((.......))).)))..((((((((((((...((((((.((((.(((.(--(........)).)))))))))))))...)))))))))))).. ( -37.30, z-score = -1.84, R) >droWil1.scaffold_181096 5725093 94 - 12416693 -----------------------ACAUUAAACGGCCCAAUACGCUCGCUGUAAAUUGUAGCAAUCAAAGCGUUGCCUUUUGCCUCACAGCUUGGUGUUACUUCUUACAGUGAGUGUG -----------------------................((((((((((((((...((((((....((((((.((.....))...)).))))..))))))...)))))))))))))) ( -30.80, z-score = -3.28, R) >droVir3.scaffold_12928 3437481 83 - 7717345 --------------------------UGAAAUGA-CGUCA-CGCCCGCUGUAAAAUGUAAUCAUCAGAGCGUC--C-UC---UCAACCGCUUGGUGUUACUUCUUACAGUGAGUGCG --------------------------........-.....-((((((((((((...((((.(((((.((((..--.-..---.....)))))))))))))...)))))))).).))) ( -25.20, z-score = -2.87, R) >droMoj3.scaffold_6328 350108 83 + 4453435 ---------------------------GAAAUGAACGUUG-CGACCGCUGUAAAUUGUAAUCAUCAAAGCGUC--U-GC---UAAAUCGCUUGGUGUUACUUCUUACAGUGAGCGCG ---------------------------............(-((..((((((((...((((.((((((.(((..--.-..---.....)))))))))))))...))))))))..))). ( -28.10, z-score = -3.51, R) >droGri2.scaffold_14853 9156922 84 - 10151454 --------------------------UUGAAUGA-CGUCA-CGCCCGCUGUAAUAUGUAAUCAUCAAAGCGUG--UAUU---UUACUCGCUUGGUGUUACUUCUUACAGUGAGUGCG --------------------------........-.....-((((((((((((...((((.((((((.(((.(--((..---.))).)))))))))))))...)))))))).).))) ( -28.30, z-score = -3.75, R) >consensus UCGCUGCAUGUGAGAGAA________UUCCGUGGCCGGCAUCGCUCGCUGUAAAUUGUAAUCAUCAAAGCGUU__UUUU__CAGAGCCGCUUGGUGUUACUUCUUACAGUGAGUGUG .........................................(((.((((((((...(((((.(((((.(((................)))))))))))))...)))))))).))).. (-22.24 = -21.32 + -0.93)

| Location | 12,956,085 – 12,956,190 |
|---|---|
| Length | 105 |
| Sequences | 11 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 70.86 |
| Shannon entropy | 0.56731 |
| G+C content | 0.46641 |
| Mean single sequence MFE | -25.05 |
| Consensus MFE | -15.25 |
| Energy contribution | -15.30 |
| Covariance contribution | 0.05 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.09 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.19 |
| SVM RNA-class probability | 0.906252 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
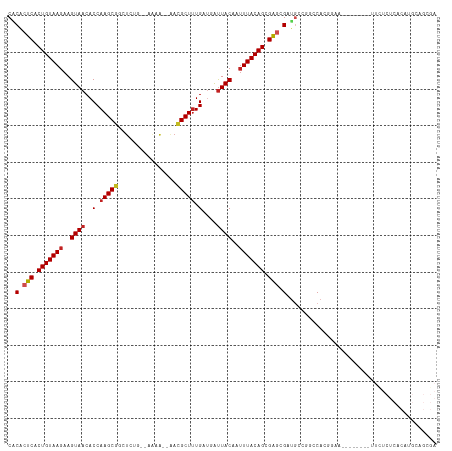
>dm3.chrX 12956085 105 - 22422827 CACACUCACUGUAAGAAGUAACACCAAGCGGCUCUG--AGAA--AACGCUUUGAUGAUUACAAUUUACAGCGAGCGAUGCCAGCCACGGAA--------UUCUCUCACAUGCAGCGA ..(.(((.(((((((..(((((((.(((((......--....--..))))).).)).))))..))))))).))).).(((..((....((.--------.....))....)).))). ( -23.80, z-score = -0.55, R) >droSec1.super_21 701213 105 - 1102487 CACACUCACUGUAAGAAGUAACACCAAGCGGCUCUG--AGAA--AACGCUUUGAUGAUUACAAUUUACAGCGAGCGAUGCCAGCCACGGAA--------UUCUCUCACAUGCAGCGA ..(.(((.(((((((..(((((((.(((((......--....--..))))).).)).))))..))))))).))).).(((..((....((.--------.....))....)).))). ( -23.80, z-score = -0.55, R) >droYak2.chrX 7229765 105 - 21770863 CACACUCACUGUAAGAAGUAACACCAAGCGGCUCUG--AGAA--AACGCUUUGAUGAUUACAAUUUACAGCGAGCGAUGCCAGCCACGGAA--------UUCUCUCACAUGCAGCGA ..(.(((.(((((((..(((((((.(((((......--....--..))))).).)).))))..))))))).))).).(((..((....((.--------.....))....)).))). ( -23.80, z-score = -0.55, R) >droEre2.scaffold_4690 4833925 105 + 18748788 CACACUCACUGUAAGAAGUAACACCAAGCGGCUCUG--AGAA--AACGCUUUGAUGAUUACAAUUUACAGCGGGCGAUGCCAGCCACGGAA--------UUCUCUCACAUGCAGCGA ..(.(((.(((((((..(((((((.(((((......--....--..))))).).)).))))..))))))).))).).(((..((....((.--------.....))....)).))). ( -22.90, z-score = 0.08, R) >droAna3.scaffold_13117 3891779 107 - 5790199 CUCACUCACUGUAAGAAGUAACACCAAGCGGCUCUGAUAAAA--AACGCUUUGAUGGUUACAAUUUACAGUGAACGAUGCCAGCCACGGAA--------UUUUCUCACAUGCAGCGA .....((((((((((..(((((.(.(((((............--..))))).)...)))))..)))))))))).((.(((.......(((.--------...))).....))).)). ( -25.74, z-score = -1.35, R) >dp4.chrXL_group3a 1299084 115 + 2690836 CACACUCACUGUAAGAAGUAACACCAAGCUGCUACCAACAAA--AAAGCUUUGGCCAUUACAAUUUACAGCGAGCGAUGGGGUCCAGGGAAAGGAGCCGCUCUCUCACAUGCUGAGA (((.(((.(((((((..((((..(((((((............--..))).))))...))))..))))))).))).).))((.(((.......))).))....(((((.....))))) ( -30.84, z-score = -0.94, R) >droPer1.super_52 413302 115 - 549863 CACACUCACUGUAAGAAGUAACACCAAGCUGCUACCAACAAA--AAAGCUUUGGCCAUUACAAUUUACAGCGAGCGAUGGGGUCCAGGGAAAGGAGCCGCUCUCUCACAUGCUGAGA (((.(((.(((((((..((((..(((((((............--..))).))))...))))..))))))).))).).))((.(((.......))).))....(((((.....))))) ( -30.84, z-score = -0.94, R) >droWil1.scaffold_181096 5725093 94 + 12416693 CACACUCACUGUAAGAAGUAACACCAAGCUGUGAGGCAAAAGGCAACGCUUUGAUUGCUACAAUUUACAGCGAGCGUAUUGGGCCGUUUAAUGU----------------------- .((.(((.(((((((..((..(((......))).((((((((((...)))))..)))))))..))))))).))).)).................----------------------- ( -21.00, z-score = 0.50, R) >droVir3.scaffold_12928 3437481 83 + 7717345 CGCACUCACUGUAAGAAGUAACACCAAGCGGUUGA---GA-G--GACGCUCUGAUGAUUACAUUUUACAGCGGGCG-UGACG-UCAUUUCA-------------------------- (((.(((.((((((((.((((((.((((((..(..---..-)--..)))).)).)).)))).)))))))).))).)-))...-........-------------------------- ( -26.90, z-score = -2.68, R) >droMoj3.scaffold_6328 350108 83 - 4453435 CGCGCUCACUGUAAGAAGUAACACCAAGCGAUUUA---GC-A--GACGCUUUGAUGAUUACAAUUUACAGCGGUCG-CAACGUUCAUUUC--------------------------- .(((..(.(((((((..(((((((.(((((.((..---..-.--))))))).).)).))))..))))))).)..))-)............--------------------------- ( -21.60, z-score = -1.88, R) >droGri2.scaffold_14853 9156922 84 + 10151454 CGCACUCACUGUAAGAAGUAACACCAAGCGAGUAA---AAUA--CACGCUUUGAUGAUUACAUAUUACAGCGGGCG-UGACG-UCAUUCAA-------------------------- (((.(((.((((((...(((((((.(((((.(((.---..))--).))))).).)).))))...)))))).))).)-))...-........-------------------------- ( -24.30, z-score = -3.17, R) >consensus CACACUCACUGUAAGAAGUAACACCAAGCGGCUCUG__AAAA__AACGCUUUGAUGAUUACAAUUUACAGCGAGCGAUGCCGGCCACGGAA________UUCUCUCACAUGCAGCGA ..(.(((.(((((((..((((..(.(((((................))))).)....))))..))))))).))).)......................................... (-15.25 = -15.30 + 0.05)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:39:01 2011