
| Sequence ID | dm3.chrX |
|---|---|
| Location | 12,935,068 – 12,935,133 |
| Length | 65 |
| Max. P | 0.817881 |

| Location | 12,935,068 – 12,935,133 |
|---|---|
| Length | 65 |
| Sequences | 3 |
| Columns | 67 |
| Reading direction | forward |
| Mean pairwise identity | 67.01 |
| Shannon entropy | 0.43859 |
| G+C content | 0.26887 |
| Mean single sequence MFE | -8.69 |
| Consensus MFE | -5.02 |
| Energy contribution | -4.58 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.70 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.79 |
| SVM RNA-class probability | 0.817881 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
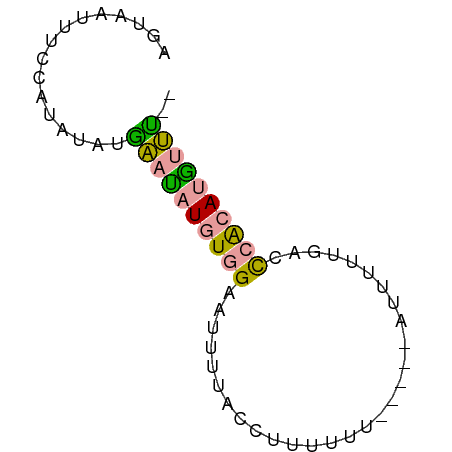
>dm3.chrX 12935068 65 + 22422827 AGUAAUUUCCAUAUAUGAAUAUGUGGAAUUUUACCUUUUUUUUCGUAUUUUUGAUCCACAUGUUC-- ................(((((((((((...............(((......))))))))))))))-- ( -12.06, z-score = -2.62, R) >droEre2.scaffold_4690 4816316 61 - 18748788 AGUAAUUUCUAUAUACAUACAUUUCGAAUACAACUUUUCU------AUUCUUGGCUAGCAAGCAUUC .........................(((((.........)------))))...(((....))).... ( -3.10, z-score = 0.50, R) >droSec1.super_21 682365 60 + 1102487 AGUAAUUUCCAUAUAUGAAUAUGUGGAAUUUUACCUUUUCU-----AUUUUUGACCCACAUAUUU-- .((((.(((((((((....)))))))))..)))).......-----...................-- ( -10.90, z-score = -2.79, R) >consensus AGUAAUUUCCAUAUAUGAAUAUGUGGAAUUUUACCUUUUUU_____AUUUUUGACCCACAUGUUU__ ................((((((((((.............................)))))))))).. ( -5.02 = -4.58 + -0.44)


| Location | 12,935,068 – 12,935,133 |
|---|---|
| Length | 65 |
| Sequences | 3 |
| Columns | 67 |
| Reading direction | reverse |
| Mean pairwise identity | 67.01 |
| Shannon entropy | 0.43859 |
| G+C content | 0.26887 |
| Mean single sequence MFE | -8.65 |
| Consensus MFE | -5.80 |
| Energy contribution | -6.03 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.55 |
| Mean z-score | -1.24 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.61 |
| SVM RNA-class probability | 0.760307 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
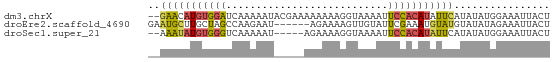
>dm3.chrX 12935068 65 - 22422827 --GAACAUGUGGAUCAAAAAUACGAAAAAAAAGGUAAAAUUCCACAUAUUCAUAUAUGGAAAUUACU --(((.(((((((.......(((..........)))....))))))).)))................ ( -10.30, z-score = -2.30, R) >droEre2.scaffold_4690 4816316 61 + 18748788 GAAUGCUUGCUAGCCAAGAAU------AGAAAAGUUGUAUUCGAAAUGUAUGUAUAUAGAAAUUACU ..((((.(((.......((((------(.(.....).))))).....))).))))............ ( -6.40, z-score = 0.10, R) >droSec1.super_21 682365 60 - 1102487 --AAAUAUGUGGGUCAAAAAU-----AGAAAAGGUAAAAUUCCACAUAUUCAUAUAUGGAAAUUACU --.((((((((((........-----..............))))))))))................. ( -9.25, z-score = -1.52, R) >consensus __AAACAUGUGGGUCAAAAAU_____AAAAAAGGUAAAAUUCCACAUAUUCAUAUAUGGAAAUUACU ..(((((((((((...........................)))))))))))................ ( -5.80 = -6.03 + 0.23)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:38:59 2011