
| Sequence ID | dm3.chrX |
|---|---|
| Location | 12,890,450 – 12,890,567 |
| Length | 117 |
| Max. P | 0.713058 |

| Location | 12,890,450 – 12,890,567 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 75.32 |
| Shannon entropy | 0.44851 |
| G+C content | 0.38666 |
| Mean single sequence MFE | -29.72 |
| Consensus MFE | -14.93 |
| Energy contribution | -18.16 |
| Covariance contribution | 3.23 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.663004 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 12890450 117 + 22422827 CUUCUGGCACCUUAUCUUUGCAUUUUUAAUUAAAUUAAAGUUGCAAACUUAAGGAGUGGAAG--GUGCACAGGAUGCUCCAAGUUUGCAAUGCGUUACCUUGGUGAACGAAACUAAGGG ......((((((((..((((((.(((((((...))))))).))))))..))))).(..((.(--(.(((.....))).))...))..)..)))....(((((((.......))))))). ( -35.20, z-score = -2.56, R) >droSim1.chrX 9819945 118 + 17042790 UUUCUAACACCUUAUCUUUGCAUUUUUAAUUAAAUUAAAGCUGCAAAGUUAAGGAGUGGGAAU-GUGCAAAGGGUGUUCAAAGUUUGCAAUGCGUUACCUUGGUGAACGAAACUAAGGG .((((.((.(((((.(((((((.(((((((...))))))).))))))).))))).)).)))).-.((((((............))))))........(((((((.......))))))). ( -34.00, z-score = -2.93, R) >droSec1.super_21 632651 118 + 1102487 UUUCUAACACCUUAUCUUUGCAUUUUUAAUUAAAUUUAAGCUGCAAAGUUACGGAGUGGGAAU-GUGCAAAGGGUGUUCAAAGUUUGCAAUGCGUUACCUUGGUGAACGAAACUAAGGG .((((.((.((.((.(((((((..(((((......))))).))))))).)).)).)).)))).-.((((((............))))))........(((((((.......))))))). ( -27.80, z-score = -0.85, R) >droYak2.chrX 7161091 118 + 21770863 UUUCUAACACCUUAUCUUUGCAUUUCUAAUUAAAUGAAAGUUACAAAGUUAAAGAGUGUGGGCAGUGCAAAGG-UGUACAAAGUUUGCACUGCGUUGCCUUGGUGAGUGAAGUGAAGGG .........((((..(((..(.((((.........)))).........(((..(((.(..(((((((((((..-(.....)..)))))))))).)..))))..))))..)))..)))). ( -32.40, z-score = -1.59, R) >droEre2.scaffold_4690 4776056 89 - 18748788 UUUCUAACACCU-AUCUUGGCAUUUCUAAUUUAAUUAAAGUGGCAAAGUUACAGAGUGGGAA--GUGCACAGGUGG--CGAGUGAAACAAAGGG------------------------- ((((...(.((.-..((((((((((((......(((...(((((...)))))..))).))))--)))).)))).))--.)...)))).......------------------------- ( -19.20, z-score = -0.58, R) >consensus UUUCUAACACCUUAUCUUUGCAUUUUUAAUUAAAUUAAAGUUGCAAAGUUAAGGAGUGGGAA__GUGCAAAGGGUGUUCAAAGUUUGCAAUGCGUUACCUUGGUGAACGAAACUAAGGG ((((((.(.(((((.(((((((.(((((((...))))))).))))))).))))).)))))))...((((((............))))))........(((((((.......))))))). (-14.93 = -18.16 + 3.23)



| Location | 12,890,450 – 12,890,567 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 75.32 |
| Shannon entropy | 0.44851 |
| G+C content | 0.38666 |
| Mean single sequence MFE | -22.49 |
| Consensus MFE | -11.84 |
| Energy contribution | -13.20 |
| Covariance contribution | 1.36 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.713058 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 12890450 117 - 22422827 CCCUUAGUUUCGUUCACCAAGGUAACGCAUUGCAAACUUGGAGCAUCCUGUGCAC--CUUCCACUCCUUAAGUUUGCAACUUUAAUUUAAUUAAAAAUGCAAAGAUAAGGUGCCAGAAG ......((((((((.((....)))))((...))......)))))...(((.((((--(((..(((.....)))(((((..((((((...))))))..)))))....))))))))))... ( -25.10, z-score = -1.23, R) >droSim1.chrX 9819945 118 - 17042790 CCCUUAGUUUCGUUCACCAAGGUAACGCAUUGCAAACUUUGAACACCCUUUGCAC-AUUCCCACUCCUUAACUUUGCAGCUUUAAUUUAAUUAAAAAUGCAAAGAUAAGGUGUUAGAAA ..........((((.((....))))))...((((((............)))))).-.(((..((.(((((.(((((((..((((((...))))))..))))))).))))).))..))). ( -26.10, z-score = -2.86, R) >droSec1.super_21 632651 118 - 1102487 CCCUUAGUUUCGUUCACCAAGGUAACGCAUUGCAAACUUUGAACACCCUUUGCAC-AUUCCCACUCCGUAACUUUGCAGCUUAAAUUUAAUUAAAAAUGCAAAGAUAAGGUGUUAGAAA .......((((...((((..((........((((((............)))))).-.........))....(((((((..((((......))))...)))))))....))))...)))) ( -21.07, z-score = -0.94, R) >droYak2.chrX 7161091 118 - 21770863 CCCUUCACUUCACUCACCAAGGCAACGCAGUGCAAACUUUGUACA-CCUUUGCACUGCCCACACUCUUUAACUUUGUAACUUUCAUUUAAUUAGAAAUGCAAAGAUAAGGUGUUAGAAA ........(((...((((..(....)((((((((((.........-..)))))))))).............(((((((..((((.........)))))))))))....))))...))). ( -28.60, z-score = -3.33, R) >droEre2.scaffold_4690 4776056 89 + 18748788 -------------------------CCCUUUGUUUCACUCG--CCACCUGUGCAC--UUCCCACUCUGUAACUUUGCCACUUUAAUUAAAUUAGAAAUGCCAAGAU-AGGUGUUAGAAA -------------------------.............((.--.((((((((((.--.........)))..(((.((...((((((...))))))...)).)))))-)))))...)).. ( -11.60, z-score = 0.23, R) >consensus CCCUUAGUUUCGUUCACCAAGGUAACGCAUUGCAAACUUUGAACACCCUUUGCAC__UUCCCACUCCUUAACUUUGCAACUUUAAUUUAAUUAAAAAUGCAAAGAUAAGGUGUUAGAAA ..............................((((((............))))))........((.(((((.(((((((..((((((...))))))..))))))).))))).))...... (-11.84 = -13.20 + 1.36)

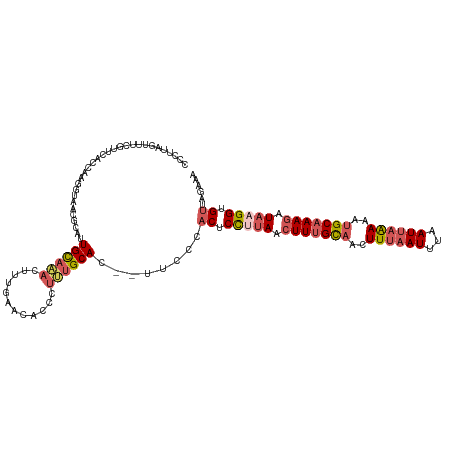

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:38:45 2011