
| Sequence ID | dm3.chrX |
|---|---|
| Location | 12,616,247 – 12,616,342 |
| Length | 95 |
| Max. P | 0.584752 |

| Location | 12,616,247 – 12,616,341 |
|---|---|
| Length | 94 |
| Sequences | 7 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 64.51 |
| Shannon entropy | 0.65860 |
| G+C content | 0.33024 |
| Mean single sequence MFE | -12.86 |
| Consensus MFE | -5.62 |
| Energy contribution | -5.36 |
| Covariance contribution | -0.26 |
| Combinations/Pair | 1.53 |
| Mean z-score | -0.90 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.584752 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
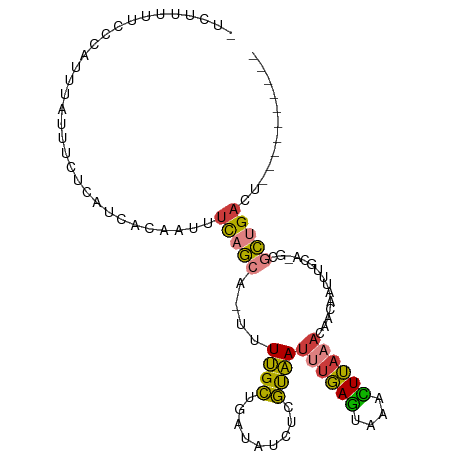
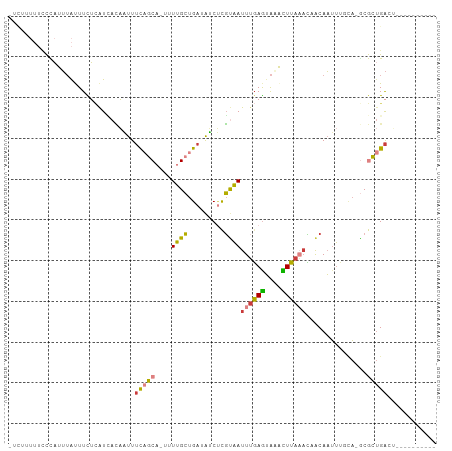
>dm3.chrX 12616247 94 - 22422827 UUCUUUUUUCCAUUUAUUUCUCAUCACAAUUUCAGCAUUUUUGCUGAUAUCUCGUAAUUUGAGUAAGCUUAAACAACAAUUUGCA-GCGCUGACU---------- ...............................(((((......((((.......((..((((((....))))))..))......))-)))))))..---------- ( -13.72, z-score = -0.75, R) >droPer1.super_9 477927 83 + 3637205 ---------UUCUUUGCUUUGUGUGAGCUUUUCCGC---UUAGAUGAUAACGUUUUAUUUUAAUACAUUAAUAAUACAAUA---AUACACUGAUGUAU------- ---------.........(..(.(((((......))---))).)..)...............((((((((...........---......))))))))------- ( -10.33, z-score = -0.18, R) >droAna3.scaffold_13117 3537506 93 - 5790199 --------UUUCUAGACUUUGGAUUUUCUUUGUCAC---UUUGCUGAUGGCCUGUAAUUGGAGAGAAUUU-UCUUGCAUAUUUUUCGCAUCGGAUCGCGCUGACU --------......(((...((.....))..)))..---....((((((.((.......)).(((((...-..........))))).))))))............ ( -12.92, z-score = 2.01, R) >droEre2.scaffold_4690 4512246 92 + 18748788 -UUCCUUUCCCAUUUAUUUCUCAUAACAAUUUCAGCA-UUUUGCUGAUAUCUCGUAAUUUGAGUAGACUUAAAUAACAAUUUCCA-ACGCUGACU---------- -...................(((........(((((.-....)))))......((.(((((((....))))))).))........-....)))..---------- ( -12.80, z-score = -2.16, R) >droYak2.chrX 6897542 93 - 21770863 -UUCCUUUCCCAUUUAUUUCUCAUCACAAUUUCAGCA-UUUUGCUGAUAUCUCGCAAUUUGAGUAGACUUAAAUAACAAUUUCUACGCGCUGACG---------- -..............................(((((.-...(((.((....)))))(((((((....)))))))..............)))))..---------- ( -13.40, z-score = -1.85, R) >droSec1.super_21 370240 94 - 1102487 UUCUUUUUCCCAUUUAUUUCUCAUCACAAUUUCAGCAUUUUUGCUGAUAUCUCGUAAUUUGAGUAAACUUAAACAACAAUUCGCA-GCGCUGACU---------- ...............................(((((......((((.......((..((((((....))))))..))......))-)))))))..---------- ( -13.42, z-score = -1.70, R) >droSim1.chrX 9637186 94 - 17042790 UUCUUUUUCCCAUUUAUUUCUCAUCACAAUUUCAGCAUUUUUGCUGAUAUCUCGUAAUUUGAGUAAACUUAAACAACAAUUCGCA-GCGCUGACU---------- ...............................(((((......((((.......((..((((((....))))))..))......))-)))))))..---------- ( -13.42, z-score = -1.70, R) >consensus _UCUUUUUCCCAUUUAUUUCUCAUCACAAUUUCAGCA_UUUUGCUGAUAUCUCGUAAUUUGAGUAAACUUAAACAACAAUUUGCA_GCGCUGACU__________ ...............................(((((....((((.........))))((((((....))))))...............)))))............ ( -5.62 = -5.36 + -0.26)

| Location | 12,616,248 – 12,616,342 |
|---|---|
| Length | 94 |
| Sequences | 7 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 64.81 |
| Shannon entropy | 0.65227 |
| G+C content | 0.32871 |
| Mean single sequence MFE | -12.70 |
| Consensus MFE | -5.62 |
| Energy contribution | -5.36 |
| Covariance contribution | -0.26 |
| Combinations/Pair | 1.53 |
| Mean z-score | -0.88 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.539332 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
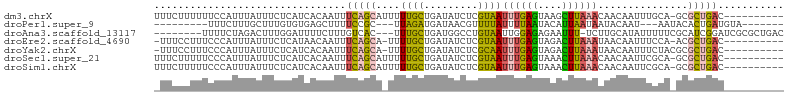
>dm3.chrX 12616248 94 - 22422827 UUUCUUUUUUCCAUUUAUUUCUCAUCACAAUUUCAGCAUUUUUGCUGAUAUCUCGUAAUUUGAGUAAGCUUAAACAACAAUUUGCA-GCGCUGAC---------- ................................(((((......((((.......((..((((((....))))))..))......))-))))))).---------- ( -13.72, z-score = -0.83, R) >droPer1.super_9 477928 83 + 3637205 ---------UUUCUUUGCUUUGUGUGAGCUUUUCCGC---UUAGAUGAUAACGUUUUAUUUUAAUACAUUAAUAAUACAAU---AAUACACUGAUGUA------- ---------..........(..(.(((((......))---))).)..)................(((((((..........---.......)))))))------- ( -9.23, z-score = 0.23, R) >droAna3.scaffold_13117 3537507 93 - 5790199 --------UUUUCUAGACUUUGGAUUUUCUUUGUCAC---UUUGCUGAUGGCCUGUAAUUGGAGAGAAUUU-UCUUGCAUAUUUUUCGCAUCGGAUCGCGCUGAC --------.......(((...((.....))..)))..---....((((((.((.......)).(((((...-..........))))).))))))........... ( -12.92, z-score = 1.88, R) >droEre2.scaffold_4690 4512247 92 + 18748788 -UUUCCUUUCCCAUUUAUUUCUCAUAACAAUUUCAGCA-UUUUGCUGAUAUCUCGUAAUUUGAGUAGACUUAAAUAACAAUUUCCA-ACGCUGAC---------- -....................(((........(((((.-....)))))......((.(((((((....))))))).))........-....))).---------- ( -12.80, z-score = -2.12, R) >droYak2.chrX 6897543 93 - 21770863 -UUUCCUUUCCCAUUUAUUUCUCAUCACAAUUUCAGCA-UUUUGCUGAUAUCUCGCAAUUUGAGUAGACUUAAAUAACAAUUUCUACGCGCUGAC---------- -...............................(((((.-...(((.((....)))))(((((((....)))))))..............))))).---------- ( -13.40, z-score = -1.86, R) >droSec1.super_21 370241 94 - 1102487 UUUCUUUUUCCCAUUUAUUUCUCAUCACAAUUUCAGCAUUUUUGCUGAUAUCUCGUAAUUUGAGUAAACUUAAACAACAAUUCGCA-GCGCUGAC---------- ................................(((((......((((.......((..((((((....))))))..))......))-))))))).---------- ( -13.42, z-score = -1.72, R) >droSim1.chrX 9637187 94 - 17042790 UUUCUUUUUCCCAUUUAUUUCUCAUCACAAUUUCAGCAUUUUUGCUGAUAUCUCGUAAUUUGAGUAAACUUAAACAACAAUUCGCA-GCGCUGAC---------- ................................(((((......((((.......((..((((((....))))))..))......))-))))))).---------- ( -13.42, z-score = -1.72, R) >consensus _UUCUUUUUCCCAUUUAUUUCUCAUCACAAUUUCAGCA_UUUUGCUGAUAUCUCGUAAUUUGAGUAAACUUAAACAACAAUUUGCA_GCGCUGAC__________ ................................(((((....((((.........))))((((((....))))))...............)))))........... ( -5.62 = -5.36 + -0.26)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:37:59 2011