
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 11,201,495 – 11,201,589 |
| Length | 94 |
| Max. P | 0.927206 |

| Location | 11,201,495 – 11,201,589 |
|---|---|
| Length | 94 |
| Sequences | 8 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 71.06 |
| Shannon entropy | 0.53533 |
| G+C content | 0.49741 |
| Mean single sequence MFE | -23.07 |
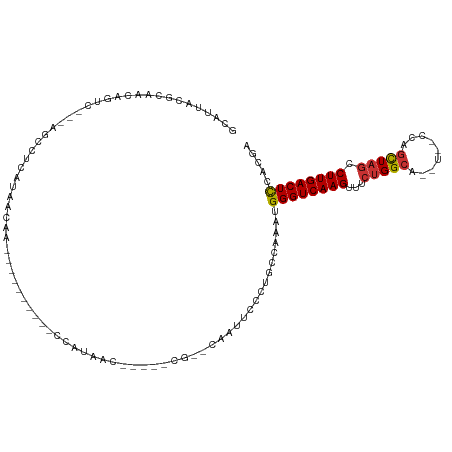
| Consensus MFE | -13.70 |
| Energy contribution | -13.93 |
| Covariance contribution | 0.22 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.39 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.33 |
| SVM RNA-class probability | 0.927206 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
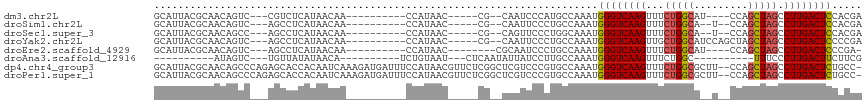
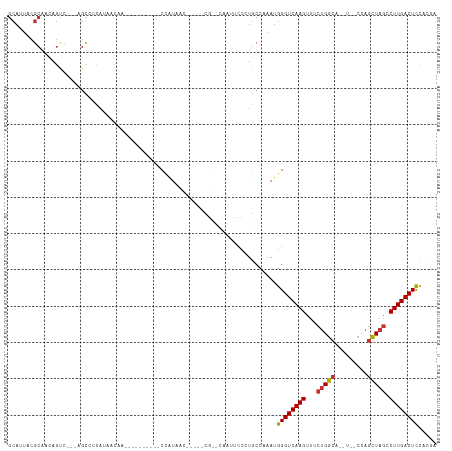
>dm3.chr2L 11201495 94 - 23011544 GCAUUACGCAACAGUC---CGUCUCAUAACAA----------CCAUAAC-----CG--CAAUCCCAUGCCAAAUGGGUCAAGUUUCUGGCAU----CCAGCUAGCCUUGACUCCACGA ((.....)).......---(((..........----------((((...-----.(--((......)))...))))((((((...(((((..----...))))).))))))...))). ( -20.30, z-score = -1.62, R) >droSim1.chr2L 11002611 94 - 22036055 GCAUUACGCAACAGUC---AGCCUCAUAACAA----------CCAUAAC-----CG--CAAUUCCCUGCCAAAUGGGUCAAGUUUCUGGCA--U--CCAGCUAGCCUUGACUCCACGA ((.....)).......---.............----------.......-----.(--((......)))....(((((((((...(((((.--.--...))))).)))))).)))... ( -19.40, z-score = -1.04, R) >droSec1.super_3 6602104 94 - 7220098 GCAUUACGCAACAGCC---AGCCUCAUAACAA----------CCAUAAC-----CG--CAGUUCCCUGGCAAAUGGGUCAAGUUUCUGGCA--U--CCAGCUAGCCUUGACUCCACGA ((.....))....(((---((...........----------.......-----..--.......)))))...(((((((((...(((((.--.--...))))).)))))).)))... ( -23.63, z-score = -1.31, R) >droYak2.chr2L 7599473 98 - 22324452 GCAUUACGCAACAGUC---AGCCUCAUAACAA----------CCAUAAC-----CG--CAAUUCCCUGCCAAAUGGGUCAAGUUGCUGGCAUCCAGCUAGCUAGCCUUGACUCCCCGA ((.....)).......---.............----------((((...-----.(--((......)))...))))((((((..((((((.....))))))....))))))....... ( -23.00, z-score = -1.06, R) >droEre2.scaffold_4929 12399297 92 + 26641161 GCAUUACGCAACAGUC---AGCCUCAUAACAA----------CCAUAAC--------CGCAAUCCCUGCCAAAUGGGUCAAGUUUCUGGCAU----CCAGCUAGCCUUGACUCCCGA- ((.....)).......---.............----------((((...--------.(((.....)))...))))((((((...(((((..----...))))).))))))......- ( -19.00, z-score = -0.92, R) >droAna3.scaffold_12916 11927500 82 + 16180835 ----------AUAGUC---UGUUAUAUAACA----------UCUGUAAU---CUCAAUAUUAUCCUUGCCAAAUGGGUCAAGUUUCUGGC----------UUUCCCUUGACUUCUUCG ----------..((((---((((....))))----------........---......................(((..((((.....))----------)).)))..))))...... ( -9.20, z-score = -0.03, R) >dp4.chr4_group3 6574301 115 - 11692001 GCAUUACGCAACAGCCCAGAGCACCACAAUCAAAGAUGAUUUCCAUAACGUUCUCGGCUCGUCCCGUGCCAAAUGGGUCAAGUUUCUGGCGCUU--CCAGCUAGCCUUGACUCUGCC- (((.((((..((((((.(((((.....(((((....)))))........))))).)))).))..))))......((((((((...(((((....--...))))).))))))))))).- ( -35.02, z-score = -2.58, R) >droPer1.super_1 8065251 115 - 10282868 GCAUUACGCAACAGCCCAGAGCACCACAAUCAAAGAUGAUUUCCAUAACGUUCUCGGCUCGUCCCGUGCCAAAUGGGUCAAGUUUCUGGCGCUU--CCAGCUAGCCUUGACUCUGCC- (((.((((..((((((.(((((.....(((((....)))))........))))).)))).))..))))......((((((((...(((((....--...))))).))))))))))).- ( -35.02, z-score = -2.58, R) >consensus GCAUUACGCAACAGUC___AGCCUCAUAACAA__________CCAUAAC_____CG__CAAUUCCCUGCCAAAUGGGUCAAGUUUCUGGCA__U__CCAGCUAGCCUUGACUCCACGA ((.....)).................................................................((((((((...(((((.........))))).))))))))..... (-13.70 = -13.93 + 0.22)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:31:46 2011