
| Sequence ID | dm3.chrX |
|---|---|
| Location | 12,515,646 – 12,515,739 |
| Length | 93 |
| Max. P | 0.602627 |

| Location | 12,515,646 – 12,515,739 |
|---|---|
| Length | 93 |
| Sequences | 8 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 69.12 |
| Shannon entropy | 0.61075 |
| G+C content | 0.44331 |
| Mean single sequence MFE | -28.26 |
| Consensus MFE | -10.90 |
| Energy contribution | -10.65 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.56 |
| Mean z-score | -1.39 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.602627 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 12515646 93 - 22422827 GGAUCUAUAUACAUAUGUAGCUGUAGACGCGAUAUCCUGCCCGU-----UGGGUGGGCAUAUGUCG-CGGGCUAUUUGCUAUAUAGCGACAAAUUGAUA ..(((........((((((((.((((.((((((((((..(((..-----.)))..))...))))))-))..))))..))))))))..........))). ( -34.07, z-score = -2.56, R) >droPer1.super_77 46337 95 + 256778 ---CGUGUAUAUAGUGGAAGAGACGGCGGCACUAGAGUACGGGUACGUGUAUAUAUGCAUAUGUCC-UGGGUUAUUUGCUAUAUAGCGACAAAUUGAUA ---(((.(((((((..((.((..(((.((((.....(((....)))(((((....))))).)))))-))..)).))..))))))))))........... ( -24.70, z-score = -0.56, R) >dp4.chrXL_group1a 3298902 95 - 9151740 ---CGUGUAUAUAGUGGAAGAGACGGCGGCACUAGAGUACGGGUACGUGUAUAUAUGCAUAUGUCC-UGGGUUAUUUGCUAUAUAGCGACAAAUUGAUA ---(((.(((((((..((.((..(((.((((.....(((....)))(((((....))))).)))))-))..)).))..))))))))))........... ( -24.70, z-score = -0.56, R) >droAna3.scaffold_13117 3411788 78 - 5790199 ------------------AUGUACAUAUGUAUC-CUUUGUAUGUAUGAGGG-AUAUGAACAUGUCG-UGACUUGUUCGCUAUAUAGCGACAAAUUGAUA ------------------(((.((((.((((((-(((..(....)..))))-)))))...))))))-)...((((.((((....))))))))....... ( -19.20, z-score = -1.24, R) >droEre2.scaffold_4690 4405330 90 + 18748788 GGGUCUAUAUACGGA--UAG--GCAGACGCGAUGUCCUGCCCGU-----UGGGUGGGCAUAUGUCGCCGGGUUAUUUGCUAUAUAGCGACAAAUUGAUA ..(.(((((((((((--(((--((.((((..((((((.((((..-----.)))))))))).)))))))....)))))).))))))))............ ( -29.10, z-score = -0.72, R) >droYak2.chrX 6792402 91 - 21770863 GAGUGUAUACGCAGU--UAG--GUAGACGCGAUAUCCUGCCUGC---UGUGGGUGGGCAUAUGUCG-CAGGUUAUUUGCUAUAUAGCGACAAAUUGAUA ..((.((..((((((--.((--((((..........))))))))---))))..)).))...(((((-(.(((.....))).....))))))........ ( -28.30, z-score = -0.89, R) >droSec1.super_21 269996 91 - 1102487 GGGUCUAUAUACAGA--UAGGUGUAGACGCGAUAUCCUGCCCGU-----UGGGUGGGCAUAUGUCG-CGGGCUAUUUGCUAUAUAGCGACAAAUUGAUA (.((((((((.....--...)))))))).).((((((..(((..-----.)))..)).))))((((-(.(((.....))).....)))))......... ( -33.00, z-score = -2.28, R) >droSim1.chrX_random 3437459 91 - 5698898 GGGUCUAUAUACAGA--UAGGUGUAGACGCGAUAUCCUGCCCGU-----UGGGUGGGCAUAUGUCG-CGGGCUAUUUGCUAUAUAGCGACAAAUUGAUA (.((((((((.....--...)))))))).).((((((..(((..-----.)))..)).))))((((-(.(((.....))).....)))))......... ( -33.00, z-score = -2.28, R) >consensus GGGUCUAUAUACAGA__UAGGUGCAGACGCGAUAUCCUGCCCGU_____UGGGUGGGCAUAUGUCG_CGGGUUAUUUGCUAUAUAGCGACAAAUUGAUA ...................................((((((((......))))))))....(((((...(((.....)))......)))))........ (-10.90 = -10.65 + -0.25)


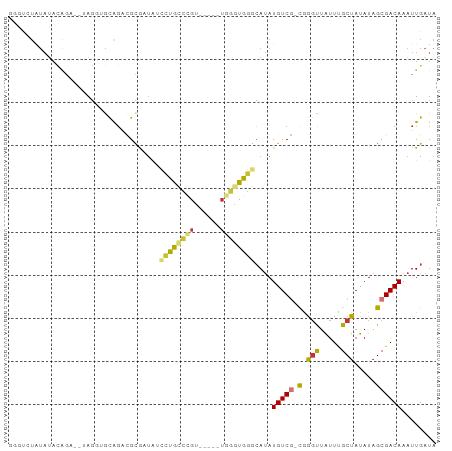
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:37:46 2011