
| Sequence ID | dm3.chrX |
|---|---|
| Location | 12,485,356 – 12,485,477 |
| Length | 121 |
| Max. P | 0.814172 |

| Location | 12,485,356 – 12,485,477 |
|---|---|
| Length | 121 |
| Sequences | 6 |
| Columns | 124 |
| Reading direction | reverse |
| Mean pairwise identity | 60.98 |
| Shannon entropy | 0.72148 |
| G+C content | 0.45044 |
| Mean single sequence MFE | -32.08 |
| Consensus MFE | -12.86 |
| Energy contribution | -13.83 |
| Covariance contribution | 0.98 |
| Combinations/Pair | 1.48 |
| Mean z-score | -1.16 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.814172 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 12485356 121 - 22422827 UCACAGCACUGUGAUAUGGUUG-GUUAAAGUAAAAGUUCAAUGGGUUAAGCAGGUGAUUAAAGGCACAUUAACCCCUAAUGGAGCUUGGACUGCACUUAACGCACGGCUCACAUGUGACCAC-- (((((....)))))..(((((.-((((((((..((((((...(((((((....(((........))).)))))))......))))))..)))....)))))(((.(.....).)))))))).-- ( -32.10, z-score = -0.41, R) >droPer1.super_12 2393986 94 + 2414086 --------------UGACUACGAGUGGCUACCAAUGCCAGUAUAUACAUGUAUGUAUUUAUGCACACACUCGAUUCCACAGAGAC----ACUAUUACCAGUAUUUCAACCAC------------ --------------......((((((((.......))).(((((((((....))))..)))))....)))))........((((.----(((......))).))))......------------ ( -19.10, z-score = -1.65, R) >droEre2.scaffold_4690 4374682 116 + 18748788 -----UCACAGUGAUAAAGUUG-GCUUAAGUAAAUGCUUGAGGGGUUGAGCAGGUGAUUAAAAGCACAUUAACCUCCCAUGGAGCUUAGGCUGCAAUUAACGCACGGCUCAAAUGUGCCCAC-- -----.................-.(((((((....)))))))(((((((((..(((.((((..(((..((((.((((...)))).))))..)))..)))))))...))))).....))))..-- ( -32.50, z-score = -0.52, R) >droYak2.chrX 6761395 106 - 21770863 ----------------UCGCUGUGAUAAAGUAAAUGCCUAAUGGGUUAAGCAGGUGAUCAAAAGCACAUUAACCUCCAUUGUUGCAUAGGCUGCAAUUAGCGCACCGCUCAAAUGUGCCCAC-- ----------------..((((......(((..((((.(((((((((((....(((........))).)))))..))))))..))))..))).....))))((((.........))))....-- ( -24.60, z-score = 0.52, R) >droSec1.super_21 239853 123 - 1102487 UCACAGCACUGUGAUAGAGUUG-GCUAAAGUAUAAGUUCAAGGGGUUAAGCAGGUGAUUAAAGGCACAUUAACCCCUAAUGGAGCUUGGACUGCACUUAACGCACUGCUCACAUGUGUGCCCAC .....((((((((((((.((((-(....(((.(((((((.(((((((((....(((........))).)))))))))....))))))).)))....)))))...))).))))).))))...... ( -40.20, z-score = -1.79, R) >droSim1.chrX 9582548 119 - 17042790 UCACAGCACUGU----GAGUUG-GCUAAAGUAAAAGUUCAAGGGGUUAAGCAGGUGAUUAAAGGCACAUUAACCCCUAAUGGAGCUUGGACUGCACUUAACGCACGGCUCACAUGUGUGCCCAC .....(((((((----((((((-.....(((..((((((.(((((((((....(((........))).)))))))))....))))))..)))((.......)).))))))))).))))...... ( -44.00, z-score = -3.13, R) >consensus _____GCACUGU__UAAAGUUG_GCUAAAGUAAAAGCUCAAGGGGUUAAGCAGGUGAUUAAAGGCACAUUAACCCCCAAUGGAGCUUGGACUGCAAUUAACGCACGGCUCACAUGUGCCCAC__ ..................((((......(((..((((((...(((((((....(((........))).)))))))......))))))..))).....))))....................... (-12.86 = -13.83 + 0.98)
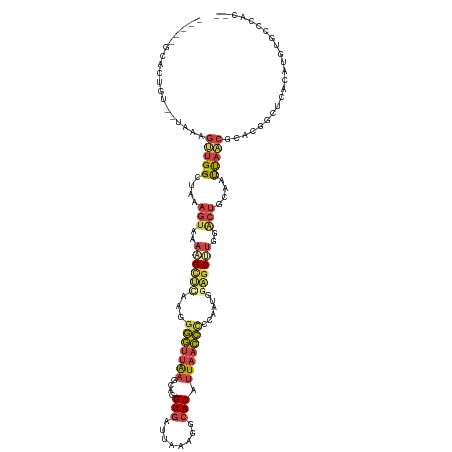


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:37:45 2011