
| Sequence ID | dm3.chrX |
|---|---|
| Location | 12,238,861 – 12,238,954 |
| Length | 93 |
| Max. P | 0.700493 |

| Location | 12,238,861 – 12,238,954 |
|---|---|
| Length | 93 |
| Sequences | 9 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 66.80 |
| Shannon entropy | 0.67701 |
| G+C content | 0.59799 |
| Mean single sequence MFE | -14.69 |
| Consensus MFE | -6.94 |
| Energy contribution | -6.62 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.25 |
| Mean z-score | -0.80 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.700493 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 12238861 93 + 22422827 CUUAAAAACCCAUUUCCAACUGUUGACACGUGCCGCAUCGUGCCGCGGCCU-UAACGCCGCCCCCGUACCCCUCUAUGCACCACCAUCCCGUUC-------- .............................(((..((((.((((.(((((..-....)))))....))))......))))..)))..........-------- ( -15.50, z-score = -0.90, R) >droPer1.super_12 2087250 83 - 2414086 CUUAAAAACCCAUUUCCAACAGUUGACACGUGCCGCAUCGUGCCGCGGCCUCGC-C--CCUUGCCCCAUGCCCCACAGUCUCAACU---------------- ....................((((((.(((((..((((.(.((.(.(((...))-)--.)..)).).))))..))).)).))))))---------------- ( -18.30, z-score = -2.26, R) >dp4.chrXL_group1a 2938522 83 + 9151740 CUUAAAAACCCAUUUCCAACAGUUGACACGUGCCGCAUCGUGCCGCGGCCUCUC-C--CCUUGCCCCAUGCCCCACAGUCUCAACU---------------- ....................((((((.(((((..((((.(.((...((......-)--)...)).).))))..))).)).))))))---------------- ( -16.60, z-score = -2.28, R) >droAna3.scaffold_13335 2217802 97 - 3335858 CUUAAAAACCCAUUUCCAACUGUCGACACGUGCCGCACCGUGCCGCGGCCUCAU-CUACCUUGCCUUCUGCCCCCUGCCCUCUCCCUCCCUUUGGCCU---- .....................(((((...(.(((((........))))))....-.......((.....))....................)))))..---- ( -11.90, z-score = 0.28, R) >droYak2.chrX 20145165 88 - 21770863 CUUAAAAACCCAUUUCCAACUGUUGACACGUACCGCGUCGUGCCGCGGCCUUAA-C---GCCACCCUGCGCCACCUCCUCUACCCCUCACUC---------- ........................(((.((...)).)))(((.((((((.....-.---))).....))).)))..................---------- ( -12.40, z-score = -0.82, R) >droEre2.scaffold_4690 4139998 87 - 18748788 CUUAAAAACCCAUUUCCAACUGUUGACACGUGCCGCAUCGUGCCGCGGCGGGACGCCGCCGUGCAACACCCCCCCCAGCCCCCCUUC--------------- ....................(((((.((((.((.((.((.(((....))).)).)).)))))))))))...................--------------- ( -24.00, z-score = -1.55, R) >droMoj3.scaffold_6359 4103509 95 - 4525533 CUUAAAAACCCAUUACCAACUGUUGACACGUGCCGCGCCGUGCCGCGACCUGUAACCC--CAAGCGCUCCCCGCCCAGAACCCCCCUCACUUACACA----- .....................((((.((((.((...))))))...)))).(((((...--...(((.....)))...((.......))..)))))..----- ( -10.90, z-score = 0.17, R) >droGri2.scaffold_15081 3900188 80 + 4274704 CUUAAAAACCCAUUUCCAACUGUUGACACGUGCCGCGUCGUGCCGCGACCUGUAACCCCUCAAGCACAC-CCCCUGCCACC--------------------- .............................((((((((......))))...((........)).))))..-...........--------------------- ( -9.70, z-score = 0.21, R) >droVir3.scaffold_12928 3869856 100 - 7717345 CUUAAAAACCCAUUUCCAACUGUUGACACGUGCCGCGUCGUGUCGCGACCUGUUACCC--CAAGCCCUCACCUCUGCCCUCGGCCUAACCCUCACCAUCACG .....................((.((((((........))))))))((...((((..(--(.((..(........)..)).))..))))..))......... ( -12.90, z-score = -0.07, R) >consensus CUUAAAAACCCAUUUCCAACUGUUGACACGUGCCGCAUCGUGCCGCGGCCUGUA_CCCCCCUACCCCACGCCCCCCAGCCCCACCUU__C____________ .....................((((.((((........))))...))))..................................................... ( -6.94 = -6.62 + -0.32)


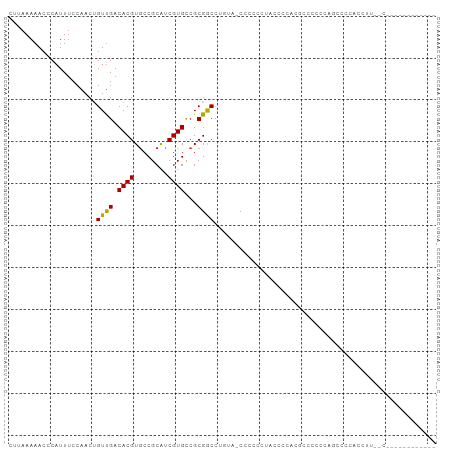
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:37:00 2011