
| Sequence ID | dm3.chrX |
|---|---|
| Location | 12,159,591 – 12,159,681 |
| Length | 90 |
| Max. P | 0.916802 |

| Location | 12,159,591 – 12,159,681 |
|---|---|
| Length | 90 |
| Sequences | 8 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 74.41 |
| Shannon entropy | 0.45196 |
| G+C content | 0.44573 |
| Mean single sequence MFE | -26.45 |
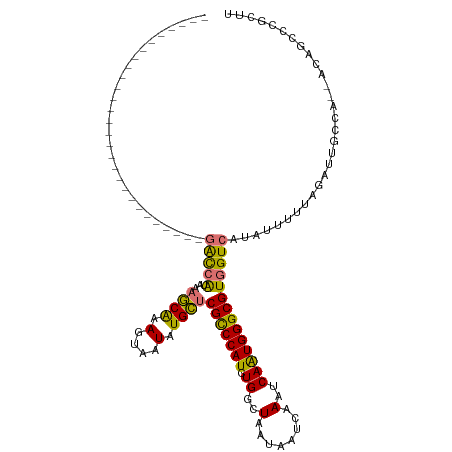
| Consensus MFE | -16.74 |
| Energy contribution | -15.90 |
| Covariance contribution | -0.84 |
| Combinations/Pair | 1.35 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.25 |
| SVM RNA-class probability | 0.916802 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
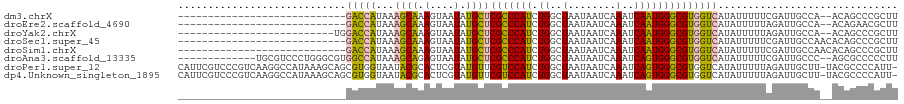
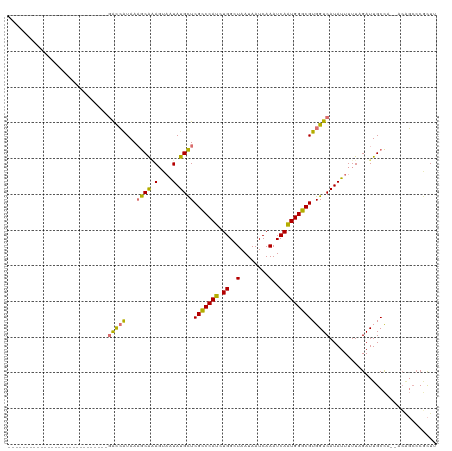
>dm3.chrX 12159591 90 - 22422827 ----------------------------GACCAUAAAGCAAAGUAAUAUGCUCGCCCAUCUGGCUAAUAAUCAAAUCAAUGGGCGUGGUCAUAUUUUUCGAUUGCCA--ACAGCCCGCUU ----------------------------((((((..((((.(....).)))).((((((.((..(........)..)))))))))))))).................--........... ( -21.90, z-score = -1.39, R) >droEre2.scaffold_4690 4065318 90 + 18748788 ----------------------------GACCAUAAAGCAAAGUAAUAUGCUCGCCCAUCUGGCUAAUAAUCAAAUCAAUGGGCGUGGUCAUAUUUUUAGAUUGCCA--ACAGAACGCUU ----------------------------((((((..((((.(....).)))).((((((.((..(........)..)))))))))))))).................--........... ( -21.90, z-score = -1.80, R) >droYak2.chrX 20069652 92 + 21770863 --------------------------UGGACCAUAAAGCAAAGUAAUAUGCUCGCCCAUCUGGCUAAUAAUCAAAUCAAUGGGCGUGGUCAUAUUUUUAGAUUGCCA--ACAGCCCGCUU --------------------------..((((((..((((.(....).)))).((((((.((..(........)..)))))))))))))).................--........... ( -21.90, z-score = -0.66, R) >droSec1.super_45 176854 92 - 245362 ----------------------------GACCAUAAAGCAAAGUAAUAUGCUCGCCCAUCUGGCUAAUAAUCAAAUCAAUGGGCGUGGUCAUAUUUUUCGAUUGCCAACACAGCCCGCUU ----------------------------((((((..((((.(....).)))).((((((.((..(........)..)))))))))))))).............................. ( -21.90, z-score = -1.38, R) >droSim1.chrX 9365471 92 - 17042790 ----------------------------GACCAUAAAGCAAAGUAAUAUGCUCGCCCAUCUGGCUAAUAAUCAAAUCAAUGGGCGUGGUCAUAUUUUUCGAUUGCCAACACAGCCCGCUU ----------------------------((((((..((((.(....).)))).((((((.((..(........)..)))))))))))))).............................. ( -21.90, z-score = -1.38, R) >droAna3.scaffold_13335 2138541 105 + 3335858 -------------UGCGUCCCUGGGCGUGGCCAUAAAGCAGAGUAAUAUGCUCGCCCAUCUGGCUAAUAAUCAAAUCAGUGGGCGUGGUCAUAUUUUUCGAUUGCCC--AGCGCCCCCUU -------------.(((...((((((((((((((......((((.....))))(((((.(((..(........)..))))))))))))))))...........))))--)))))...... ( -38.50, z-score = -2.47, R) >droPer1.super_12 1995238 118 + 2414086 CAUUCGUCCCGUCAAGGCCAUAAAGCAGCGUGGUAAUACGCACUCGUAUGUUCGUCCAUCUGGCUAAUAAUCAAAUCAGUGGGCGUGGUCAUAUUUUUAGAUUGCUU-UACGCCCCAUU- ...............(((..((((((((((((....)))))....(((((..((((((.(((..(........)..)))))))))....)))))........)))))-)).))).....- ( -31.80, z-score = -2.02, R) >dp4.Unknown_singleton_1895 44965 118 + 56477 CAUUCGUCCCGUCAAGGCCAUAAAGCAGCGUGGUAAUACGCACUCGUAUGUUCGUCCAUCUGGCUAAUAAUCAAAUCAGUGGGCGUGGUCAUAUUUUUAGAUUGCUU-UACGCCCCAUU- ...............(((..((((((((((((....)))))....(((((..((((((.(((..(........)..)))))))))....)))))........)))))-)).))).....- ( -31.80, z-score = -2.02, R) >consensus ____________________________GACCAUAAAGCAAAGUAAUAUGCUCGCCCAUCUGGCUAAUAAUCAAAUCAAUGGGCGUGGUCAUAUUUUUAGAUUGCCA__ACAGCCCGCUU ............................(((((...((((.(....).))))(((((((.((..(........)..)))))))))))))).............................. (-16.74 = -15.90 + -0.84)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:36:50 2011