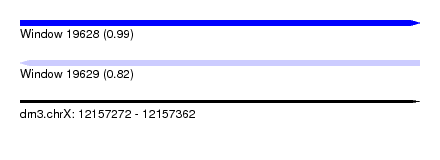

| Sequence ID | dm3.chrX |
|---|---|
| Location | 12,157,272 – 12,157,362 |
| Length | 90 |
| Max. P | 0.994316 |

| Location | 12,157,272 – 12,157,362 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 83.21 |
| Shannon entropy | 0.24911 |
| G+C content | 0.47444 |
| Mean single sequence MFE | -30.55 |
| Consensus MFE | -20.93 |
| Energy contribution | -21.74 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.08 |
| Mean z-score | -3.36 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.69 |
| SVM RNA-class probability | 0.994316 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
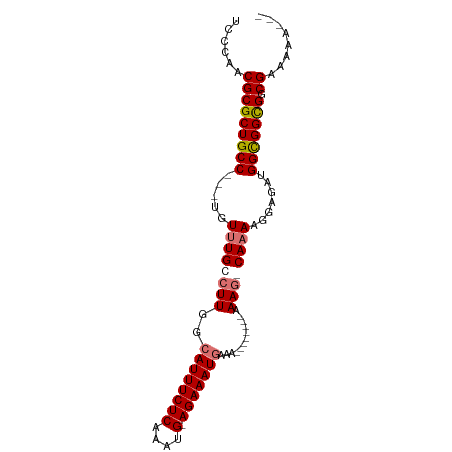
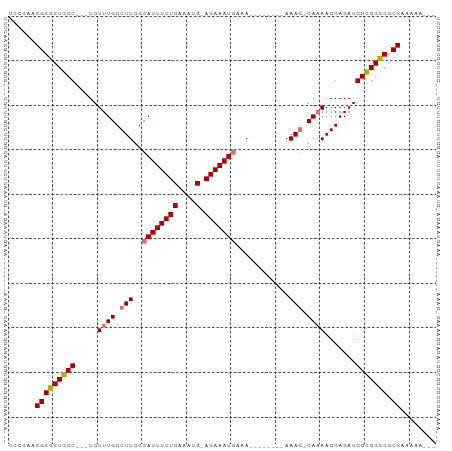
>dm3.chrX 12157272 90 + 22422827 UCCCAACGCGCUGCC---UGUUUGCCUUGGCAUUUCUCAAACGGAGAAAUAAAACAAAAAAAAAAAGCAAAAGGAGAUGGUGGCGACGAAAAG--- ......(((((..((---.((((.((((.(((((((((.....)))))))................))..))))))))))..))).)).....--- ( -23.49, z-score = -1.97, R) >droSim1.chrX 9363220 81 + 17042790 UCCCAACGCGCUGCC---UGUUUGCCUUGGCAUUUCUCAAAUG-AGAAAUGAAA--------AAAGGCACAAGGAGAUGGCGGCGGCGAAAAG--- ......(((((((((---(.((((((((..((((((((....)-)))))))...--------.))))))....)).).)))))).))).....--- ( -33.40, z-score = -3.90, R) >droYak2.chrX 20067448 83 - 21770863 UCCCAACGCGCUGCC---UGUUUGCCUUGGCAUUUCUCAAAUG-AGAAAUGAAA--------AAAG-CAAAAGGAGAUGGCGGUGGCGGUAAAAAG ......(((((((((---..(((((.((..((((((((....)-)))))))...--------)).)-)))).......)))))).)))........ ( -28.50, z-score = -2.77, R) >droEre2.scaffold_4690 4063111 86 - 18748788 UCCCAACGCGCUGCCGCCUGUUUGCCUUGGCAUUUCUCAAAUG-AGAAAUGAAA--------AAAG-CAAAAGGAGAUGGUGGCGGCGUUGAAAAG ...((((((((..((.(((.(((((.((..((((((((....)-)))))))...--------)).)-)))))))....))..)).))))))..... ( -36.80, z-score = -4.80, R) >consensus UCCCAACGCGCUGCC___UGUUUGCCUUGGCAUUUCUCAAAUG_AGAAAUGAAA________AAAG_CAAAAGGAGAUGGCGGCGGCGAAAAA___ ......(((((((((.....((((((((..((((((((....).)))))))............)))))))).......))))))).))........ (-20.93 = -21.74 + 0.81)

| Location | 12,157,272 – 12,157,362 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 83.21 |
| Shannon entropy | 0.24911 |
| G+C content | 0.47444 |
| Mean single sequence MFE | -27.36 |
| Consensus MFE | -14.39 |
| Energy contribution | -15.20 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.78 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.79 |
| SVM RNA-class probability | 0.819353 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
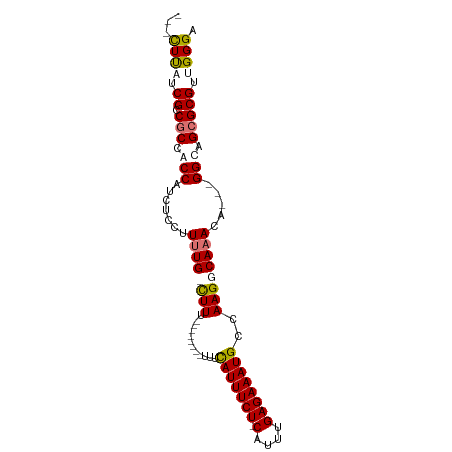
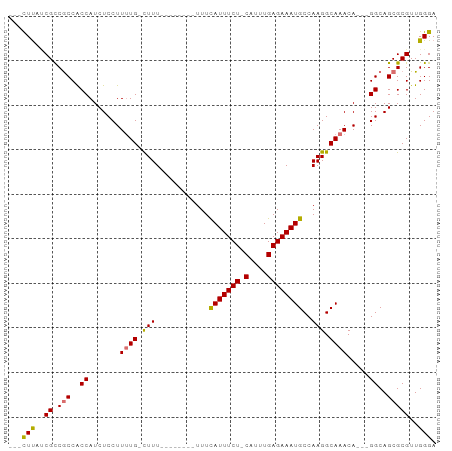
>dm3.chrX 12157272 90 - 22422827 ---CUUUUCGUCGCCACCAUCUCCUUUUGCUUUUUUUUUUUGUUUUAUUUCUCCGUUUGAGAAAUGCCAAGGCAAACA---GGCAGCGCGUUGGGA ---(((..((.(((.............(((((.(((.(((((...((((((((.....)))))))).))))).))).)---)))))))))..))). ( -18.81, z-score = -0.09, R) >droSim1.chrX 9363220 81 - 17042790 ---CUUUUCGCCGCCGCCAUCUCCUUGUGCCUUU--------UUUCAUUUCU-CAUUUGAGAAAUGCCAAGGCAAACA---GGCAGCGCGUUGGGA ---(((..(((.((.(((.........((((((.--------...(((((((-(....))))))))..))))))....---))).)))))..))). ( -28.82, z-score = -2.81, R) >droYak2.chrX 20067448 83 + 21770863 CUUUUUACCGCCACCGCCAUCUCCUUUUG-CUUU--------UUUCAUUUCU-CAUUUGAGAAAUGCCAAGGCAAACA---GGCAGCGCGUUGGGA ..........(((.(((...(((((((((-((((--------...(((((((-(....))))))))..)))))))).)---)).)).))).))).. ( -28.10, z-score = -3.73, R) >droEre2.scaffold_4690 4063111 86 + 18748788 CUUUUCAACGCCGCCACCAUCUCCUUUUG-CUUU--------UUUCAUUUCU-CAUUUGAGAAAUGCCAAGGCAAACAGGCGGCAGCGCGUUGGGA ..(..((((((.((..((....(((((((-((((--------...(((((((-(....))))))))..)))))))).))).))..))))))))..) ( -33.70, z-score = -4.48, R) >consensus ___CUUAUCGCCGCCACCAUCUCCUUUUG_CUUU________UUUCAUUUCU_CAUUUGAGAAAUGCCAAGGCAAACA___GGCAGCGCGUUGGGA ...((((.((.(((.(((............................((((((.(....)))))))((....))........))).))))).)))). (-14.39 = -15.20 + 0.81)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:36:49 2011