
| Sequence ID | dm3.chrX |
|---|---|
| Location | 11,401,917 – 11,402,034 |
| Length | 117 |
| Max. P | 0.694823 |

| Location | 11,401,917 – 11,402,034 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 86.15 |
| Shannon entropy | 0.21359 |
| G+C content | 0.33678 |
| Mean single sequence MFE | -25.20 |
| Consensus MFE | -18.15 |
| Energy contribution | -17.77 |
| Covariance contribution | -0.37 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.72 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.678752 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
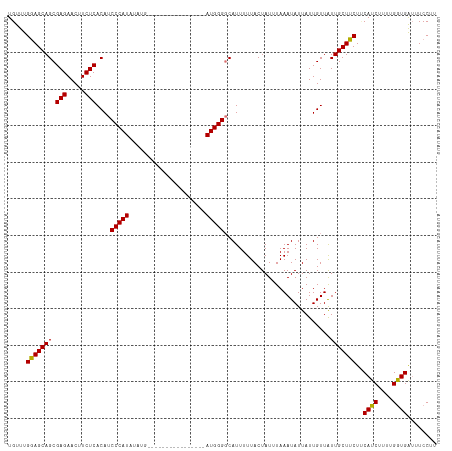
>dm3.chrX 11401917 117 + 22422827 UGUUUGGAGCAGCGAGAACUUCUCACAUCCCAUAUAUGUAUGUAUGUACAUAUGAUGGGGCAUUUUUACUAUUUAAAUAUUAUUGUUAUUGCUUCUUCAUCUUUUGGUGAUUUCCUU .....(((((((((((.....)))....(((((((((((((....)))))))).)))))))..............((((....))))..)))))).(((((....)))))....... ( -30.60, z-score = -3.08, R) >droSec1.super_36 123767 117 + 449511 UGUUUGGAGCAGCGAGAACUUCUCACAUCCCAUAUAUGCAUGAAUGUACAUAUGAUGGGGCAUUUUUACUAUAUAAAUAUUAUUGUUGUUGCUUCUUCAUCUUUCGGUGGAUUCCUU .....(((((((((((.....)))...(((((((((((.((....)).))))).))))))...........................))))))))((((((....))))))...... ( -28.70, z-score = -1.53, R) >droYak2.chrX 11016482 101 - 21770863 UGUUUGGAGCAGCGAGAACUUCUCACAUCCCAUAUAUG----------------AUGGGCCAUUUUUACUAUUUAAAUAUUAUUGUUAUUGCUUCUACAUCUUUUGAUGAUUUCCUU .....(((((((.(((.....)))....(((((.....----------------))))).............................)))))))..((((....))))........ ( -19.80, z-score = -1.66, R) >droEre2.scaffold_4690 14993560 101 - 18748788 UGUUUGAAGCAGCGAGAACUUCUCACAUCCCAUAUAUG----------------AUGGGGCAUUUUUACUAUUUAAAUAUUAUUGUUAUUGCUUCUCCAUCUUUUGGUGAUUUCCUU .....(((((((((((.....)))....(((((.....----------------)))))))..............((((....))))..))))))..((((....))))........ ( -21.70, z-score = -1.74, R) >consensus UGUUUGGAGCAGCGAGAACUUCUCACAUCCCAUAUAUG________________AUGGGGCAUUUUUACUAUUUAAAUAUUAUUGUUAUUGCUUCUUCAUCUUUUGGUGAUUUCCUU .....(((((((.(((.....)))....(((((.....................))))).............................)))))))..((((....))))........ (-18.15 = -17.77 + -0.37)


| Location | 11,401,917 – 11,402,034 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 86.15 |
| Shannon entropy | 0.21359 |
| G+C content | 0.33678 |
| Mean single sequence MFE | -23.30 |
| Consensus MFE | -15.20 |
| Energy contribution | -17.70 |
| Covariance contribution | 2.50 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.65 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.694823 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
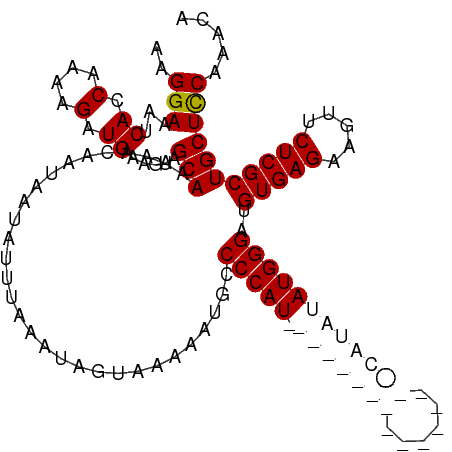
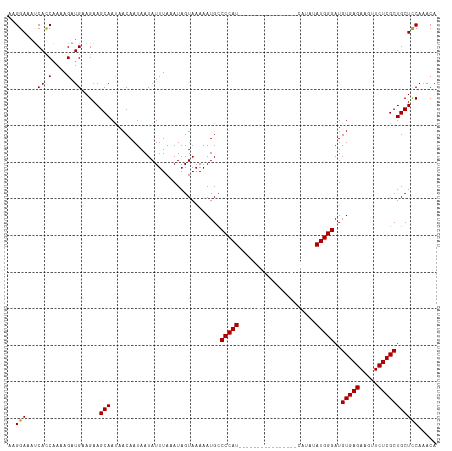
>dm3.chrX 11401917 117 - 22422827 AAGGAAAUCACCAAAAGAUGAAGAAGCAAUAACAAUAAUAUUUAAAUAGUAAAAAUGCCCCAUCAUAUGUACAUACAUACAUAUAUGGGAUGUGAGAAGUUCUCGCUGCUCCAAACA ..(((..(((.(....).)))....(((..............................(((((.(((((((......))))))))))))..(((((.....)))))))))))..... ( -24.60, z-score = -2.82, R) >droSec1.super_36 123767 117 - 449511 AAGGAAUCCACCGAAAGAUGAAGAAGCAACAACAAUAAUAUUUAUAUAGUAAAAAUGCCCCAUCAUAUGUACAUUCAUGCAUAUAUGGGAUGUGAGAAGUUCUCGCUGCUCCAAACA ..(((...((.(....).)).....(((..............................(((((.(((((((......))))))))))))..(((((.....)))))))))))..... ( -26.70, z-score = -2.41, R) >droYak2.chrX 11016482 101 + 21770863 AAGGAAAUCAUCAAAAGAUGUAGAAGCAAUAACAAUAAUAUUUAAAUAGUAAAAAUGGCCCAU----------------CAUAUAUGGGAUGUGAGAAGUUCUCGCUGCUCCAAACA ..(((...((((....)))).....(((..............................(((((----------------.....)))))..(((((.....)))))))))))..... ( -19.90, z-score = -1.66, R) >droEre2.scaffold_4690 14993560 101 + 18748788 AAGGAAAUCACCAAAAGAUGGAGAAGCAAUAACAAUAAUAUUUAAAUAGUAAAAAUGCCCCAU----------------CAUAUAUGGGAUGUGAGAAGUUCUCGCUGCUUCAAACA ..........(((.....))).((((((..............................(((((----------------.....)))))..(((((.....)))))))))))..... ( -22.00, z-score = -2.30, R) >consensus AAGGAAAUCACCAAAAGAUGAAGAAGCAAUAACAAUAAUAUUUAAAUAGUAAAAAUGCCCCAU________________CAUAUAUGGGAUGUGAGAAGUUCUCGCUGCUCCAAACA ..(((...((.(....).)).....(((..............................(((((.(((((((......))))))))))))..(((((.....)))))))))))..... (-15.20 = -17.70 + 2.50)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:34:25 2011