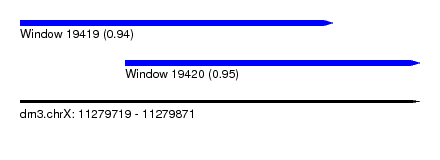

| Sequence ID | dm3.chrX |
|---|---|
| Location | 11,279,719 – 11,279,871 |
| Length | 152 |
| Max. P | 0.952187 |

| Location | 11,279,719 – 11,279,838 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 95.80 |
| Shannon entropy | 0.06611 |
| G+C content | 0.42437 |
| Mean single sequence MFE | -26.95 |
| Consensus MFE | -26.21 |
| Energy contribution | -26.52 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.97 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.41 |
| SVM RNA-class probability | 0.936733 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 11279719 119 + 22422827 CCCACGUGCCCAACAGCAAUCGCUCACCAAUUAAAAGCACCACCAGGUGGCGCAAAUUAAAAGUGAUUAAACCUGCACAGCAGUUAAUUGAGCCAUUACUUUAAUUAAUUGCUGGAAUU .((..((((......((....))((((...((((..((.((((...)))).))...))))..))))........))))((((((((((((((.......)))))))))))))))).... ( -30.50, z-score = -2.56, R) >droEre2.scaffold_4690 14873703 119 - 18748788 CCCACGUGCCCAACAGCAAUCGCUCACCAAUUAAAAGCACCACCAGGUGGCACAAAUGAAAAGUGAUUAAGCCCGCACAGCCGUUAAUUGAGCCAUUACUUUAAUUAAUUGCUGGAAUU .....(((((....(((....)))..............(((....)))))))).........(((........))).((((.((((((((((.......)))))))))).))))..... ( -26.10, z-score = -1.13, R) >droYak2.chrX 10897246 119 - 21770863 CCCACGUGCCAAACAGCAAUCGCUCACCAAUUAAAAGCACCACCAGGUGGCACAAAUGAAAAGUGAUUAAAACCGCACAGCAGUUAAUUGAGCCAUUACUUUAAUUAAUUACUGGAAUU .....(((((....(((....)))..............(((....)))))))).........(((........))).(((.(((((((((((.......))))))))))).)))..... ( -24.20, z-score = -1.48, R) >droSim1.chrX 8709801 119 + 17042790 CCCACGUGCCCAACAGCAAUCGCUCACCAAUUAAAAGCACCACCAGGUGGCGCAAAUUAAAAGUGAUUAAAACCGCACAGCAGUUAAUUGAGCCAUUACUUUAAUUAAUCGCUGGAAUU .((..((((......((....))((((...((((..((.((((...)))).))...))))..))))........))))(((.((((((((((.......)))))))))).))))).... ( -27.00, z-score = -1.95, R) >consensus CCCACGUGCCCAACAGCAAUCGCUCACCAAUUAAAAGCACCACCAGGUGGCACAAAUGAAAAGUGAUUAAAACCGCACAGCAGUUAAUUGAGCCAUUACUUUAAUUAAUUGCUGGAAUU .....(((((....(((....)))..............(((....)))))))).........(((........))).(((((((((((((((.......)))))))))))))))..... (-26.21 = -26.52 + 0.31)



| Location | 11,279,759 – 11,279,871 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 92.55 |
| Shannon entropy | 0.11767 |
| G+C content | 0.39110 |
| Mean single sequence MFE | -28.20 |
| Consensus MFE | -25.14 |
| Energy contribution | -25.70 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.89 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.58 |
| SVM RNA-class probability | 0.952187 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 11279759 112 + 22422827 CACCAGGUGGCGCAAAUUAAAAGUGAUUAAACCUGCACAGCAGUUAAUUGAGCCAUUACUUUAAUUAAUUGCUGGAAUUCUUAAUAAAUUCACAGCCAUACGGUUGACACGU ...(((((..(((.........))).....)))))..(((((((((((((((.......)))))))))))))))(((((.......))))).(((((....)))))...... ( -30.30, z-score = -3.12, R) >droEre2.scaffold_4690 14873743 110 - 18748788 CACCAGGUGGCACAAAUGAAAAGUGAUUAAGCCCGCACAGCCGUUAAUUGAGCCAUUACUUUAAUUAAUUGCUGGAAUUUGUGAUA--UUCACAGCCAUACGGUUGACACGU ......(((.(((((((.....(((........))).((((.((((((((((.......)))))))))).))))..)))))))...--....(((((....))))).))).. ( -29.40, z-score = -1.93, R) >droYak2.chrX 10897286 111 - 21770863 CACCAGGUGGCACAAAUGAAAAGUGAUUAAAACCGCACAGCAGUUAAUUGAGCCAUUACUUUAAUUAAUUACUGGAAUUUGUGAUAG-UUCACAGCCAUACGGUUGACACAU .(((..(((((...........(((........))).(((.(((((((((((.......))))))))))).))).....(((((...-.))))))))))..)))........ ( -26.90, z-score = -2.17, R) >droSim1.chrX 8709841 112 + 17042790 CACCAGGUGGCGCAAAUUAAAAGUGAUUAAAACCGCACAGCAGUUAAUUGAGCCAUUACUUUAAUUAAUCGCUGGAAUUCGUAAUAAAUUCACAGCCAUACGGUUGACACGU .(((..(((((...........(((........))).((((.((((((((((.......)))))))))).))))(((((.......)))))...)))))..)))........ ( -26.20, z-score = -1.83, R) >consensus CACCAGGUGGCACAAAUGAAAAGUGAUUAAAACCGCACAGCAGUUAAUUGAGCCAUUACUUUAAUUAAUUGCUGGAAUUCGUAAUAA_UUCACAGCCAUACGGUUGACACGU .(((..(((((...........(((........))).(((((((((((((((.......)))))))))))))))....................)))))..)))........ (-25.14 = -25.70 + 0.56)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:33:59 2011