
| Sequence ID | dm3.chrX |
|---|---|
| Location | 11,067,003 – 11,067,099 |
| Length | 96 |
| Max. P | 0.634267 |

| Location | 11,067,003 – 11,067,097 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 60.08 |
| Shannon entropy | 0.67177 |
| G+C content | 0.45015 |
| Mean single sequence MFE | -20.34 |
| Consensus MFE | -8.96 |
| Energy contribution | -7.44 |
| Covariance contribution | -1.52 |
| Combinations/Pair | 1.93 |
| Mean z-score | -0.90 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.634267 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
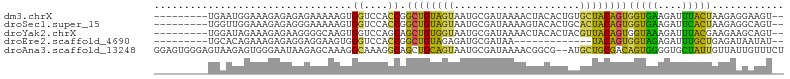
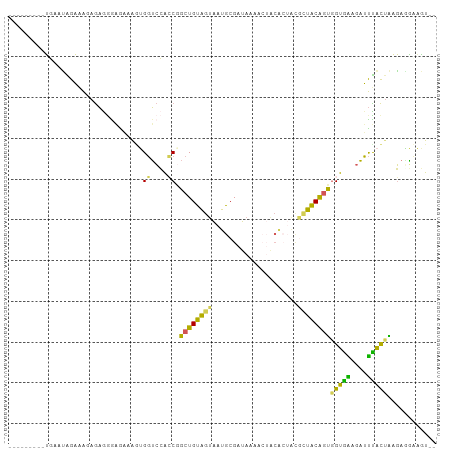
>dm3.chrX 11067003 94 - 22422827 ---------UGAAUGGAAAGAGAGAGAAAAAGUGGUCCACCGGCUGUAGUAAUGCGAUAAAACUACACUGUGCUACAGUGGUGAAGAUUUACUAAGAGGAAGU-- ---------.....................((((((((((((.((((((((.((...........))...)))))))))))))..)).)))))..........-- ( -18.10, z-score = -0.88, R) >droSec1.super_15 1770722 94 - 1954846 ---------UGGUUGGAAAGAGAGGGAAAAAGUGGUCCACCGGCUGUAGUAAUGCGAUAAAAGUACACUGCACUACAGUGGUGAAGAUUCACUAAGAGGCAGU-- ---------.((((((........(((........))).))))))(((....)))...........(((((.((....((((((....))))))..)))))))-- ( -22.00, z-score = -0.72, R) >droYak2.chrX 10709870 94 + 21770863 ---------UGGAUAGAAAGAGAAGGGGCAAGUGGUCCAGCAGCUGUGGUAAUGCGAUAAAACUACACUACGUUACAGUGGUAAAGAUUUACGAAGAAGCAGU-- ---------............(...((((.....))))..).(((((.......((.((((.((..(((((......)))))..)).)))))).....)))))-- ( -18.10, z-score = -0.34, R) >droEre2.scaffold_4690 14684519 81 + 18748788 ---------UGCACAGAAAGAGAGGAGGAAGUGGGUCCACCGGCUGUAGAGAUGCGAUAA-------------UACAGUGGUAGAGAUUUGCUGAGAUAAUAU-- ---------....................((..(((((((((.(((((............-------------))))))))).).))))..))..........-- ( -16.30, z-score = -1.22, R) >droAna3.scaffold_13248 2613798 103 - 4840945 GGAGUGGGAGUAAGAGUGGGAAUAAGAGCAAAGGCAAAGGCAGCUGCAGUAAUGCGAUAAAACGGCG--AUGCUGCGACAGUGGGGUGCUAUUGUUAUUGUUUCU .................((((((((.(((((.((((...((.(.(((((((.(((.........)))--.))))))).).))....)))).))))).)))))))) ( -27.20, z-score = -1.37, R) >consensus _________UGAAUAGAAAGAGAGGGAGAAAGUGGUCCACCGGCUGUAGUAAUGCGAUAAAACUACACUACGCUACAGUGGUGAAGAUUUACUAAGAGGAAGU__ .................................((....)).((((((((.....................))))))))(((((....)))))............ ( -8.96 = -7.44 + -1.52)

| Location | 11,067,005 – 11,067,099 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 74.82 |
| Shannon entropy | 0.41518 |
| G+C content | 0.44441 |
| Mean single sequence MFE | -14.67 |
| Consensus MFE | -8.26 |
| Energy contribution | -9.58 |
| Covariance contribution | 1.31 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.00 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>dm3.chrX 11067005 94 + 22422827 UUCCUCUUAGUAAAUCUUCACCACUGUAGCACAGUGUAGUUUUAUCGCAUUACUACAGCCGGUGGACCACUUUUUCUCUCUCUUUCCAUUCACA ................((((((.((((((...((((..((...))..)))).))))))..))))))............................ ( -16.00, z-score = -2.39, R) >droSec1.super_15 1770724 94 + 1954846 UGCCUCUUAGUGAAUCUUCACCACUGUAGUGCAGUGUACUUUUAUCGCAUUACUACAGCCGGUGGACCACUUUUUCCCUCUCUUUCCAACCACA ........((((....((((((.((((((((..(((.........)))..))))))))..)))))).))))....................... ( -20.60, z-score = -2.44, R) >droYak2.chrX 10709872 94 - 21770863 UGCUUCUUCGUAAAUCUUUACCACUGUAACGUAGUGUAGUUUUAUCGCAUUACCACAGCUGCUGGACCACUUGCCCCUUCUCUUUCUAUCCACA .........((((...((((.((((((...((((((..((...))..)))))).)))).)).))))....)))).................... ( -10.90, z-score = 0.27, R) >droEre2.scaffold_4690 14684521 81 - 18748788 AUUAUCUCAGCAAAUCUCUACCACUGUA-------------UUAUCGCAUCUCUACAGCCGGUGGACCCACUUCCUCCUCUCUUUCUGUGCACA .........(((....((((((.(((((-------------............)))))..))))))......................)))... ( -11.17, z-score = -1.25, R) >consensus UGCCUCUUAGUAAAUCUUCACCACUGUAGCGCAGUGUAGUUUUAUCGCAUUACUACAGCCGGUGGACCACUUUCCCCCUCUCUUUCCAUCCACA ................((((((.((((((....(((.........)))....))))))..))))))............................ ( -8.26 = -9.58 + 1.31)
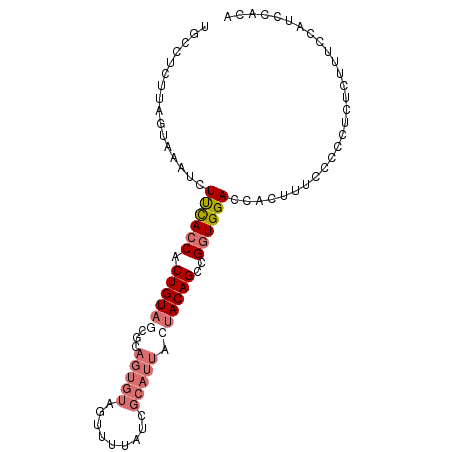



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:33:25 2011