| Sequence ID | dm3.chrX |
|---|---|
| Location | 10,942,335 – 10,942,482 |
| Length | 147 |
| Max. P | 0.736925 |

| Location | 10,942,335 – 10,942,482 |
|---|---|
| Length | 147 |
| Sequences | 5 |
| Columns | 147 |
| Reading direction | reverse |
| Mean pairwise identity | 69.73 |
| Shannon entropy | 0.49909 |
| G+C content | 0.51895 |
| Mean single sequence MFE | -44.72 |
| Consensus MFE | -20.40 |
| Energy contribution | -20.64 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.736925 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
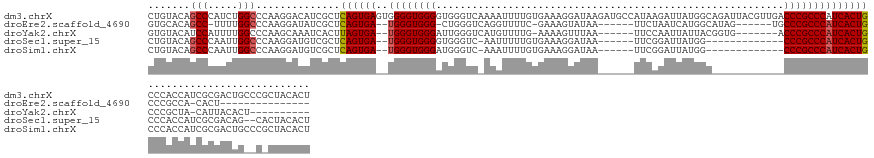
>dm3.chrX 10942335 147 - 22422827 CUGUACAGCCCAUCUGGCCCAAGGACAUCGCUCAGUGAGUGGGGUGGGGUGGGUCAAAAUUUUGUGAAAGGAUAAGAUGCCAUAAGAUUAUGGCAGAUUACGUUGACCCGCCCAUCACUGCCCACCAUCGCGACUGCCCGCUACACU .(((((.(((.....)))....((.((((((...(((.(((((((((((((((((((.(((((((......)))))))((((((....))))))........)))))))))))..)))..)))))))).)))).)).))).)))).. ( -56.30, z-score = -2.44, R) >droEre2.scaffold_4690 14567117 114 + 18748788 GUGCACAGCC-UUUUGGCCCAAGGAUAUCGCUCAGUGA--UGGGUGGG-CUGGGUCAGGUUUUC-GAAAGUAUAA------UUCUAAUCAUGGCAUAG------UGCCCGCCCAUCACUGCCCGCCA-CACU--------------- (((....(((-....)))....((....((..((((((--((((((((-(...((((((((...-(((.......------))).)))).))))....------.)))))))))))))))..)))).-))).--------------- ( -45.20, z-score = -2.64, R) >droYak2.chrX 10595241 120 + 21770863 GUGUACAUCCAUUUUGGCCCAAGCAAAUCACUUAGUGA--UGGGUGGGAUUGGGUCAUGUUUUG-AAAAGUUUAA------UUCCAAUUAUUACGGUG-------ACCCGCCCAUCACUGCCCGCUA-CAUUACACU---------- (((((...........((....))........((((((--((((((((((((((...((..((.-...))..)).------.))))))).....(...-------.)))))))))))))).......-...))))).---------- ( -36.10, z-score = -1.90, R) >droSec1.super_15 1654090 123 - 1954846 CUGUACAGCCCAAUUGGCCCAAGGAUGUCGCUCAGUGA--UGGGUGGGGUGGGUC-AAUUUUUGUGAAAGGAUAA------UUCGGAUUAUGG-------------CCCGCCCAUCACUGCCCACCAUCGCGACAG--CACUACACU .(((...(((.....)))...((..((((((...(((.--(((((.(((((((((-(.....(.((((.......------)))).)...)))-------------)))))))......))))).))).)))))).--..))))).. ( -42.00, z-score = -0.56, R) >droSim1.chrX 8496632 125 - 17042790 CUGUACAGCCCAAUUGGCCCAAGGAUGUCGCUCAGUGA--UGGGUGGGAUGGGUC-AAAUUUUGUGAAAGGAUAA------UUCGGAUUAUGG-------------CCCGCCCAUCACUGCCCACCAUCGCGACUGCCCGCUACACU .(((((.(((.....)))....((..(((((.((((((--((((((((...((((-.....((((......))))------....))))....-------------)))))))))))))).........)))))...))).)))).. ( -44.00, z-score = -1.00, R) >consensus CUGUACAGCCCAUUUGGCCCAAGGAUAUCGCUCAGUGA__UGGGUGGGAUGGGUC_AAAUUUUGUGAAAGGAUAA______UUCGGAUUAUGGC____________CCCGCCCAUCACUGCCCACCAUCGCGACAG__C_CUACACU .......(((.....)))..............((((((..((((((((..........................................................))))))))))))))........................... (-20.40 = -20.64 + 0.24)

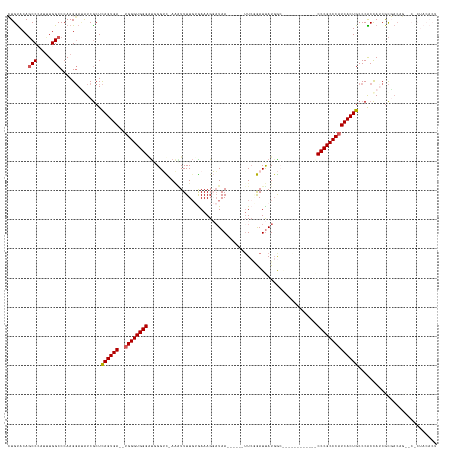
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:33:07 2011