
| Sequence ID | dm3.chrX |
|---|---|
| Location | 10,804,930 – 10,805,033 |
| Length | 103 |
| Max. P | 0.828166 |

| Location | 10,804,930 – 10,805,033 |
|---|---|
| Length | 103 |
| Sequences | 12 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 84.41 |
| Shannon entropy | 0.31747 |
| G+C content | 0.46685 |
| Mean single sequence MFE | -26.11 |
| Consensus MFE | -20.80 |
| Energy contribution | -20.81 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.80 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.82 |
| SVM RNA-class probability | 0.828166 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 10804930 103 + 22422827 -------CUCGAC-ACUCGUAUCGAUCCUGAGCGGCCCUCGAGUUGCGUUUAAAUUUCAAUUUGCUGCCACAUAAGCGACACUUGAGUGCGCCGCUUUUU-UGCUUAUCUAC-- -------.((((.-.......))))....(((((((((((((((..((((((.....((......)).....))))))..))))))).).)))))))...-...........-- ( -28.10, z-score = -1.99, R) >droSim1.chrX 8367088 103 + 17042790 -------CUCGAC-ACUCGUAUCGAUCCUGAGCGGCCCUCGAGUUGCGUUUAAAUUUCAAUUUGCUGCCACAUAAGCGACACUUGAGUGCGCCGCUUUUU-UGCUUAUCUAC-- -------.((((.-.......))))....(((((((((((((((..((((((.....((......)).....))))))..))))))).).)))))))...-...........-- ( -28.10, z-score = -1.99, R) >droSec1.super_15 1508214 103 + 1954846 -------CUCGAC-ACUCGUAUCGAUCCUGAGCGGCCCUCGAGUUGCGUUUAAGUUUCAAUUUGCUGCCACAUAAGCGACACUUGAGUGCGCCGCUUUUU-UGCUUAUCUAC-- -------.((((.-.......))))....(((((((((((((((..((((((.((..((......))..)).))))))..))))))).).)))))))...-...........-- ( -30.50, z-score = -2.28, R) >droYak2.chrX 10455993 103 - 21770863 -------CUCGAC-ACUCGUAUCGAUCCUGAGCGGCCCUCGAGUUGCGUUUAAAUUUCAAUUUGCUGCCACAUAAGCGACACUCGAGUGCGCCGCUUUUU-UGCUUAUCUAC-- -------.((((.-.......))))....(((((((((((((((..((((((.....((......)).....))))))..))))))).).)))))))...-...........-- ( -30.20, z-score = -2.48, R) >droEre2.scaffold_4690 14426610 102 - 18748788 -------CUCGAC-ACUCGUAUCGAUCCUGAGCGGCCCUCGAGUUGCGUUUAAAUUUCAAUUUGCUGCCACAUAAGCGACACUUGAGUGCGCCGCUUUUU--GCUUAUCUAC-- -------.((((.-.......))))....(((((((((((((((..((((((.....((......)).....))))))..))))))).).)))))))...--..........-- ( -28.10, z-score = -1.95, R) >droAna3.scaffold_13417 2323409 107 + 6960332 -------CUCACCCACUCGUAUCGAUCCUGAGCAGCCCUCGAGCUGCGUUUAAAUUUCAAUUUGCUGCCACAUAAGCGACACUUGAGUGCGCCGCUUUUUGCGCUUAUAUAUCC -------................(((..(((((.((.((((((.((((((((.....((......)).....)))))).)))))))).))...((.....)))))))...))). ( -23.80, z-score = -0.97, R) >dp4.chrXL_group1e 622723 110 + 12523060 GUGUACAUGCACAUAC-CGUAUCGAUCCUGAGCGGCCCUCGAGUUGCGUUUAAAUUUCAAUUUGCUGCCACAUAAGCGACACUUAAGUGCACCGCUUUUU-UGCUUAUCUUC-- ((((....))))....-...........((((((((...(((((((.(.......).)))))))..)))....(((((.(((....)))...)))))...-.))))).....-- ( -23.00, z-score = -0.39, R) >droPer1.super_30 370098 110 - 958394 AUGUACAUGCACAUAC-CGUAUCGAUCCUGAGCGGCCCUCGAGUUGCGUUUAAAUUUCAAUUUGCUGCCACAUAAGCGACACUUAAGUGCACCGCUUUUU-UGCUUAUCUUC-- ((((......))))..-...........((((((((...(((((((.(.......).)))))))..)))....(((((.(((....)))...)))))...-.))))).....-- ( -20.10, z-score = 0.36, R) >droWil1.scaffold_180702 858417 103 + 4511350 -------CUUGAG-ACACAUAUCGAUCCUGAGCGCUACUCAAGCAGCGUUUAAAUUUCAAUUUGCUGCCACAUAAACGUCACUUGAGUGCACUGCUUUUU-UCUUUUUUUGU-- -------......-.........((....((((((.(((((((..(((((((.....((......)).....)))))))..)))))))))...))))...-)).........-- ( -21.10, z-score = -0.78, R) >droVir3.scaffold_12928 398190 100 + 7717345 ------UUUUGCC--UUUGCAUCGAUCCUAAGCGGCCCUCAAGCUGCGUUUAAAUUUCAAUUUUCUGCCACAUGAGCGUCACUUGAGUGCGCCGCUUUUU--GCUUAUCU---- ------.......--...(((........((((((((((((((..(((((((.....((......)).....)))))))..)))))).).)))))))..)--))......---- ( -28.00, z-score = -2.45, R) >droMoj3.scaffold_6328 3999085 103 - 4453435 ------UUUUGCC--UUUGCAUCGAUUCUAAGCGGCCCUCGAGCUGCGUUUAAAUUUCAAUUUUCUGCCACAUGAGCGUCACUUGAGUGCGCCUCUUUUU-UGCUUAUCUGC-- ------.......--...(((........(((.((((((((((..(((((((.....((......)).....)))))))..)))))).).))).)))...-)))........-- ( -22.30, z-score = -0.13, R) >droGri2.scaffold_15203 6830831 102 + 11997470 ------UUUUGCC--UUUGCAUCGAACCUAAGCGGCACUCAAGCAGCGUUUAAAUUUCAAUUUUCUGACACAUGAGCGUCACUUGAGUGCGCCGCUUUUU--GCUUAUCUAC-- ------.......--...(((........((((((((((((((..(((((((....(((......)))....)))))))..)))))))..)))))))..)--))........-- ( -30.00, z-score = -2.87, R) >consensus _______CUCGAC_ACUCGUAUCGAUCCUGAGCGGCCCUCGAGUUGCGUUUAAAUUUCAAUUUGCUGCCACAUAAGCGACACUUGAGUGCGCCGCUUUUU_UGCUUAUCUAC__ .............................((((((((((((((.((((((((.....((......)).....)))))).)))))))).).)))))))................. (-20.80 = -20.81 + 0.01)


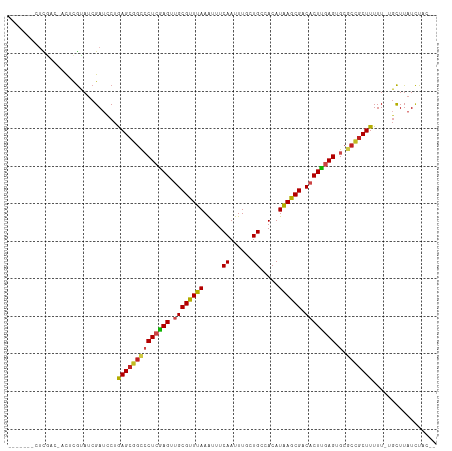
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:32:34 2011