
| Sequence ID | dm3.chrX |
|---|---|
| Location | 10,706,805 – 10,706,964 |
| Length | 159 |
| Max. P | 0.724194 |

| Location | 10,706,805 – 10,706,964 |
|---|---|
| Length | 159 |
| Sequences | 3 |
| Columns | 162 |
| Reading direction | forward |
| Mean pairwise identity | 69.43 |
| Shannon entropy | 0.39679 |
| G+C content | 0.45759 |
| Mean single sequence MFE | -44.97 |
| Consensus MFE | -23.18 |
| Energy contribution | -24.30 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.724194 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
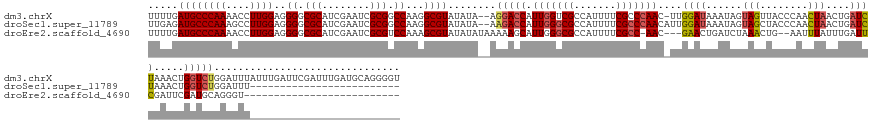
>dm3.chrX 10706805 159 + 22422827 UUUUGAUGCCCAAAACCUUGGAGGGGCGCAUCGAAUCGCGGCCAAGGCGUAUAUA--AGGACCAUUGGUCGCCAUUUUCGCCCAAC-UUGGAUAAAUAGUAGUUACCCAACUAACUGAUCUAAACUGGUCUGGAUUUAUUUGAUUCGAUUUGAUGCAGGGGU .......((((....(((....)))..(((((((((((((((((((((.(.....--).).)).)))))))).......(.(((..-((((((...((((((((....)))).))))))))))..))).).(((((.....)))))))))))))))..)))) ( -49.60, z-score = -1.48, R) >droSec1.super_11789 1094 135 + 1231 UUGAGAUGCCCAAAGCCUUGGAGGGGCGCAUCGAAUCGCGGCCAAGGCGUAUAUA--AAGACCAUUGGGCGCCAUUUUCGCCCAACAUUGGAUAAAUAGUAGCUACCCAACUAACUGAUCUAAACUGGUCUGGAUUU------------------------- .........((...(((((((....(((........)))..))))))).......--.(((((((((((((.......)))))))..((((((...((((.........))))....))))))..))))))))....------------------------- ( -40.90, z-score = -1.06, R) >droEre2.scaffold_4690 14325285 130 - 18748788 UUUUGAUGCCCAAAACCUUGGAGGGGCGCAUCGAAUCGCGUCCAAAGCGUAUAUAUAAAAAGCAUUGGGCGCCAUUUUCGCC-AAC---GAACUGAUCUAAACUG--AAUUUAUUUGAUUCGAUUCGAUGCAGGGU-------------------------- .......((((....(((....)))..((((((((((((((((((.((.............)).))))))))....((((..-..)---)))............(--((((.....))))))))))))))).))))-------------------------- ( -44.42, z-score = -3.29, R) >consensus UUUUGAUGCCCAAAACCUUGGAGGGGCGCAUCGAAUCGCGGCCAAGGCGUAUAUA__AAGACCAUUGGGCGCCAUUUUCGCCCAAC_UUGGAUAAAUAGUAGCUACCCAACUAACUGAUCUAAACUGGUCUGGAUUU_________________________ .....((((((((....))))..((.(((........))).))...))))........(((((.(((((((.......)))))))....((((......(((........)))....)))).....)))))............................... (-23.18 = -24.30 + 1.12)


| Location | 10,706,805 – 10,706,964 |
|---|---|
| Length | 159 |
| Sequences | 3 |
| Columns | 162 |
| Reading direction | reverse |
| Mean pairwise identity | 69.43 |
| Shannon entropy | 0.39679 |
| G+C content | 0.45759 |
| Mean single sequence MFE | -40.77 |
| Consensus MFE | -27.53 |
| Energy contribution | -28.10 |
| Covariance contribution | 0.57 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.25 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.629731 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 10706805 159 - 22422827 ACCCCUGCAUCAAAUCGAAUCAAAUAAAUCCAGACCAGUUUAGAUCAGUUAGUUGGGUAACUACUAUUUAUCCAA-GUUGGGCGAAAAUGGCGACCAAUGGUCCU--UAUAUACGCCUUGGCCGCGAUUCGAUGCGCCCCUCCAAGGUUUUGGGCAUCAAAA ......(((((.(((((...............((((................((((((((.......))))))))-(((((.((.......)).)))))))))..--.......((....))..))))).)))))((((..(....)....))))....... ( -39.70, z-score = -0.05, R) >droSec1.super_11789 1094 135 - 1231 -------------------------AAAUCCAGACCAGUUUAGAUCAGUUAGUUGGGUAGCUACUAUUUAUCCAAUGUUGGGCGAAAAUGGCGCCCAAUGGUCUU--UAUAUACGCCUUGGCCGCGAUUCGAUGCGCCCCUCCAAGGCUUUGGGCAUCUCAA -------------------------...((((((...((..(((((.....(((((((((.......)))))))))((((((((.......))))))))))))).--.))....(((((((..(((........)))....)))))))))))))........ ( -44.90, z-score = -1.88, R) >droEre2.scaffold_4690 14325285 130 + 18748788 --------------------------ACCCUGCAUCGAAUCGAAUCAAAUAAAUU--CAGUUUAGAUCAGUUC---GUU-GGCGAAAAUGGCGCCCAAUGCUUUUUAUAUAUACGCUUUGGACGCGAUUCGAUGCGCCCCUCCAAGGUUUUGGGCAUCAAAA --------------------------.....((((((((((((((.......)))--)............(((---(..-..))))....(((.((((.((.............)).)))).)))))))))))))((((..(....)....))))....... ( -37.72, z-score = -1.82, R) >consensus _________________________AAAUCCAGACCAGUUUAGAUCAGUUAGUUGGGUAGCUACUAUUUAUCCAA_GUUGGGCGAAAAUGGCGCCCAAUGGUCUU__UAUAUACGCCUUGGCCGCGAUUCGAUGCGCCCCUCCAAGGUUUUGGGCAUCAAAA ............................((((((........((((.......(((((((.......)))))))..((((((((.......))))))))))))...........(((((((..(((........)))....)))))))))))))........ (-27.53 = -28.10 + 0.57)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:32:20 2011