| Sequence ID | dm3.chrX |
|---|---|
| Location | 10,567,429 – 10,567,489 |
| Length | 60 |
| Max. P | 0.954994 |

| Location | 10,567,429 – 10,567,489 |
|---|---|
| Length | 60 |
| Sequences | 5 |
| Columns | 67 |
| Reading direction | reverse |
| Mean pairwise identity | 63.28 |
| Shannon entropy | 0.60998 |
| G+C content | 0.32191 |
| Mean single sequence MFE | -7.20 |
| Consensus MFE | -4.70 |
| Energy contribution | -4.38 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.00 |
| Structure conservation index | 0.65 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.61 |
| SVM RNA-class probability | 0.954994 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
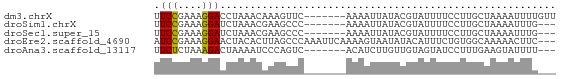
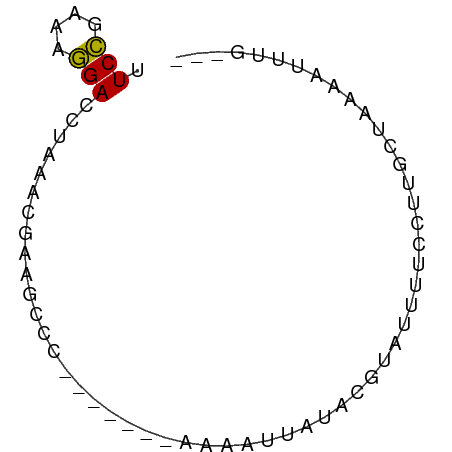
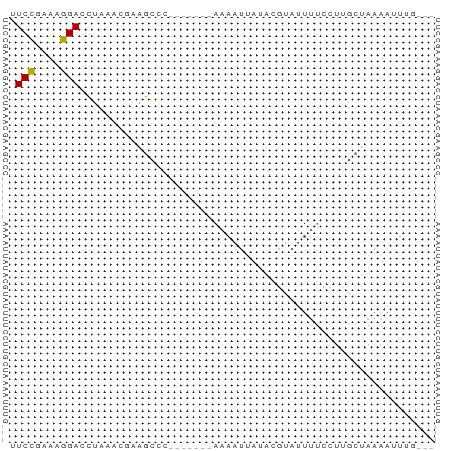
>dm3.chrX 10567429 60 - 22422827 UUCCGAAAGGACCUAAACAAAGUUC-------AAAAUUAUACGUAUUUUCCUUGCUAAAAUUUUGUU .(((....))).............(-------((((((.((.(((.......))))).))))))).. ( -9.60, z-score = -2.09, R) >droSim1.chrX 8199438 57 - 17042790 UUCCGAAAGGAUCUAAACGAAGCCC-------AAAAUUAUACGUAUUUUCCUUGCUAAAAUUUG--- .(((....))).........(((..-------(((((.......)))))....)))........--- ( -6.60, z-score = -0.92, R) >droSec1.super_15 1271804 57 - 1954846 UUCCGAAAGGAUCUAAACGAAGCCC-------AAAAUUAUACGUAUUUUCCUUGCUAAAAUUUG--- .(((....))).........(((..-------(((((.......)))))....)))........--- ( -6.60, z-score = -0.92, R) >droEre2.scaffold_4690 14191744 64 + 18748788 AUCCGAAAGGAACUACACUUAGCCCAAAUUCAAAAGUAAUAUACAUUUCUGUGGCAAAAACUUC--- .(((....)))..........(((((.......................)).))).........--- ( -9.20, z-score = -1.48, R) >droAna3.scaffold_13117 1206849 57 + 5790199 UUCUCUAAAGACUAAAAUCCCAGUC-------ACAUCUUGUUGUAGUAUCCUUUGAAGUAUUUU--- (((......((((........))))-------(((......)))..........))).......--- ( -4.00, z-score = 0.41, R) >consensus UUCCGAAAGGACCUAAACGAAGCCC_______AAAAUUAUACGUAUUUUCCUUGCUAAAAUUUG___ .(((....)))........................................................ ( -4.70 = -4.38 + -0.32)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:31:55 2011