
| Sequence ID | dm3.chrX |
|---|---|
| Location | 10,360,421 – 10,360,516 |
| Length | 95 |
| Max. P | 0.559403 |

| Location | 10,360,421 – 10,360,516 |
|---|---|
| Length | 95 |
| Sequences | 13 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 73.95 |
| Shannon entropy | 0.53768 |
| G+C content | 0.63640 |
| Mean single sequence MFE | -36.12 |
| Consensus MFE | -13.02 |
| Energy contribution | -13.47 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.44 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.559403 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 10360421 95 - 22422827 GGGGUAAUCUGUACGCUCCAUUCCAUUCGCUCGAUCAUUCGAGCCGUUCGC---CCGCCUACUCGGCG------GCGGCGGCCGCC---------CACAGCUACCCGAACCCU-- .(((((..((((..((............((((((....)))))).(..(((---(((((.....))))------..))))..))).---------.)))).))))).......-- ( -37.30, z-score = -2.35, R) >droSim1.chrX_random 2936360 95 - 5698898 GGGGUAAUCUGUACGCUCCAUUCCAUUCGCUCGAUCAUUCGAGCCGUUCGC---CCGCCUACUCGGCG------GCGGCGGCCGCC---------CACAGCUUCCAGAACCCU-- (((((..((((...(((...........((((((....))))))((..(((---(((((.....))))------..))))..))..---------...)))...)))))))))-- ( -37.10, z-score = -2.45, R) >droSec1.super_15 1067562 95 - 1954846 GGGGUAAUCUGUACGCUCCAUUCCAUUCGCUCGAUCAUUCGAGCCGUUCGC---CCGCCUACUCGGCG------GCGGCGGCCGCC---------CACAGCUUUCAGAACCCU-- (((((..((((...(((...........((((((....))))))((..(((---(((((.....))))------..))))..))..---------...)))...)))))))))-- ( -37.80, z-score = -2.78, R) >droYak2.chrX 18933956 95 - 21770863 GGGGUAAUCUGUACGCUCCAUUCCAUUCGCUCGAUCAUUCGAGCCGUUCGC---CCGCCUACUCGGCG------GCGGCGGCCGCC---------CACAGCUUCCCGAACCAC-- ((((....((((..((............((((((....)))))).(..(((---(((((.....))))------..))))..))).---------.))))..)))).......-- ( -35.30, z-score = -1.86, R) >droEre2.scaffold_4690 13980523 95 + 18748788 GGGGUAAUCUGUACGCUCCAUUCCAUUCGCUCGAUCAUUCGAGCCGUUCGC---CCGCCUACUCGGCG------GCGGCGGCCGCC---------CACAGCUUCCCGAACCAC-- ((((....((((..((............((((((....)))))).(..(((---(((((.....))))------..))))..))).---------.))))..)))).......-- ( -35.30, z-score = -1.86, R) >droAna3.scaffold_13335 983185 101 + 3335858 GCGGCAAUCUGUACGCUCCAUUCCAUUCGCUCGAUCAUUCGAGCCGCUCGCCAGCUGCAUACUCGGCA------GCGGCGGCGGCC------GCCCACAGCUUCCCGGGCCAC-- (((((.......................((((((....))))))(((.((((.(((((.......)))------))))))))))))------)).....(((.....)))...-- ( -42.30, z-score = -2.33, R) >dp4.chrXL_group1e 5706598 95 - 12523060 GCGGCAAUCUGUAUGCUCCAUUCCAUACGCUCGAUCAUUCGAGCCGCUCGC---CCGCCUAUUCGGCG------GCAGCGGCCGCA---------CACAGCUUCCCGGGCCAC-- (((((.....(((((........)))))((((((....))))))((((.((---(.(((.....))))------))))))))))).---------....(((.....)))...-- ( -40.80, z-score = -3.44, R) >droPer1.super_21 634320 95 - 1645244 GCGGCAAUCUGUAUGCUCCAUUCCAUACGCUCGAUCAUUCGAGCCGCUCGC---CCGCCUAUUCGGCG------GCAGCGGCCGCA---------CACAGCUUCCCGGGCCAC-- (((((.....(((((........)))))((((((....))))))((((.((---(.(((.....))))------))))))))))).---------....(((.....)))...-- ( -40.80, z-score = -3.44, R) >droWil1.scaffold_180702 878895 107 + 4511350 GUGCCAAUCUGUA---UUCAUUCCAUUCGAUUGAACAUUCGAGCACGCCGG---CCGCCUAUUCUGCAGCCGUUGCGGCAGCUGCAUCAGCCGCUCACAAUUUUCCGGGCCAU-- (.(((........---((((.((.....)).)))).....((((.((((((---(.((.......)).))))..)))...((((...)))).))))...........))))..-- ( -29.20, z-score = 0.04, R) >droVir3.scaffold_12932 222151 104 - 2102469 CGGGCAAUCUGUACGCUCCAUUCCAUUCGCUCGAUCAUUCGAACCGCUCGC---CCGCCUACUCAGCG------GCAGCAGCUGCUGCCGCUGCGCACAGUUUCCCGGGCCAU-- .((((.........((............))((((....))))...))))((---(((...((((((((------(((((....)))))))))).....)))....)))))...-- ( -40.20, z-score = -2.48, R) >droMoj3.scaffold_6473 14873867 98 + 16943266 CUGGCAAUCUGUACGCUUCAUUCCAUUCGCUUGAUCAUUCGAACCGCUCGC---CCGCAUACUCAGCG------GCCGCCGCCGCG------GCCCAUUGCCUGCCCGGCCAC-- .((((.....(((.((............((((((....))))...((....---..)).......))(------(((((....)))------)))....)).)))...)))).-- ( -25.90, z-score = 0.64, R) >droGri2.scaffold_15203 4655079 104 + 11997470 CGGGCAAUCUGUACGCUCCAUUCCAUUCGCUUGAUCAUUCGAGCCGCUCAC---CCGCCUAUUCAGCG------GCAGCGGCCGCUGCCGCGGCGCACAGUUUUCCGGGUCAU-- ((((.((.((((.((((...........((((((....)))))).((....---..)).......(((------(((((....)))))))))))).)))))).))))......-- ( -40.30, z-score = -1.91, R) >anoGam1.chr2R 60086852 87 + 62725911 ------GGCGUCAUGCAUCAGUUCAAGCAGUUGACGGAUCCAUCGAUAC-----CGAUCUGUGCGGCG------CCGUCGCCCA-----------AAUCGACCGGCGGAACGCAG ------.(((((.((((.(((.((((....))))((((((....))).)-----))..))))))).((------((((((....-----------...))).)))))).)))).. ( -27.30, z-score = 0.29, R) >consensus GGGGCAAUCUGUACGCUCCAUUCCAUUCGCUCGAUCAUUCGAGCCGCUCGC___CCGCCUACUCGGCG______GCGGCGGCCGCC_________CACAGCUUCCCGGGCCAC__ ..(((...((((................((((((....))))))...........(((.......)))......))))..)))................................ (-13.02 = -13.47 + 0.45)


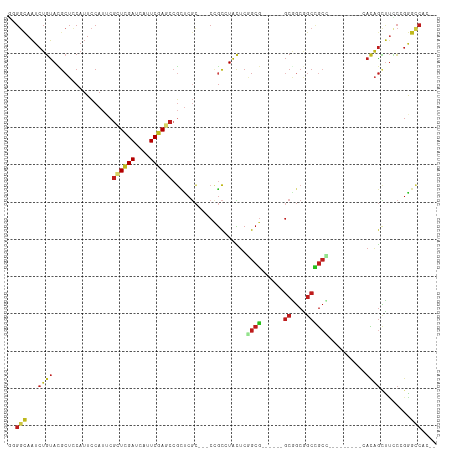
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:31:21 2011