
| Sequence ID | dm3.chrX |
|---|---|
| Location | 10,304,036 – 10,304,176 |
| Length | 140 |
| Max. P | 0.685990 |

| Location | 10,304,036 – 10,304,176 |
|---|---|
| Length | 140 |
| Sequences | 5 |
| Columns | 165 |
| Reading direction | reverse |
| Mean pairwise identity | 74.80 |
| Shannon entropy | 0.40506 |
| G+C content | 0.42831 |
| Mean single sequence MFE | -43.97 |
| Consensus MFE | -23.58 |
| Energy contribution | -24.78 |
| Covariance contribution | 1.20 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.685990 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 10304036 140 - 22422827 AGGUUUUACAGGCUCUGUUGGGCCCUGUUGAUUAACAUGGCAUGUAGAAGCAAGUUCGUGGCCACAGAAAGCUUUUCGGAGGUAAUGGUUAAGAAAAGCUAUGGUAAUAGAGCCAG------GAACCUGGUAGCUUUCUAAAGCUU------------------- (((((((...(((((((((((((((((((....)))).(((.(((....))).))).).)))).((...((((((((...............)))))))).)).)))))))))).)------))))))...(((((....))))).------------------- ( -41.76, z-score = -0.64, R) >droEre2.scaffold_4690 13919833 154 + 18748788 ACAUUCUAUGGGC---UUUUAG--CUGUUGGUUAACAUGGCAUGUAGAAGCAAGUGCUUGGCCACAGAAAGCUUUUCGGAGUAAGUGGUUAAGAAAAGCUAUGGUAAUAGAGCCUG------GAAGCUGGCGGCUUUCUGAAGCUCUGGUUUGAACAAUAGUUAC .((..(((.((((---(((.((--((((..(((..((.(((.(.((..(.(((((.(((((((((.....((((....))))..)))))))))....))).)).)..)).))))))------..)))..))))))....))))))))))..))............ ( -50.80, z-score = -2.05, R) >droYak2.chrX 18876829 155 - 21770863 AGAUUUAACUGUU---UUUUGGCCCUGUUAAUUAACAUGGCAUG------UAUGUU-ACAGCCACAGAAAGCUUUUCGGAGUUGAUGGUUCAGAAAAGCUACGGUAAUGGAGCCUGGUAAUAGAACCUGGCAGCUUUCAAAAGCUCUGGUUGAAACAAUAGUUAC .....(((((((.---.....(((.((((....)))).)))...------..((((-.((((((.....(((((((..((((((..(((((......((((.(((......)))))))....)))))...))))))..))))))).)))))).))))))))))). ( -48.50, z-score = -2.86, R) >droSec1.super_15 1000139 132 - 1954846 AGAUUUCACAGGC---AUCGGACCCUGUUGAUUAACAUGGCAAGU-----UAAGUUCUUGGCCACAGGAAGCUUUUCGGAGUUAAUGGUUAAGAAAAGCUAUGGUAAUAGAGCCAG------GAAACUGGCAGCUUUCUAAAGCUC------------------- .....(((((((.---.......)))))..((((.((((((....-----....((((((((((.....(((((....)))))..))))))))))..)))))).)))).))(((((------....)))))(((((....))))).------------------- ( -40.10, z-score = -1.87, R) >droSim1.chrX 8023796 132 - 17042790 AGAUUUCACAGGC---AUCGGACCCUGUUGAUUAACAUGGCAUGU-----UAAGUUCUUGGCCACAGGAAGCUUUUCGGAGUUAAUGGCUAAGAAAAGCUAUGGUAAUAGAGCCAG------GAAACUGACAGCUUUCUAAAGCUC------------------- .....(((((((.---.......)))).)))(((((((...))))-----)))(((((..((((.....((((((((..(((.....)))..)))))))).))))...)))))(((------....)))..(((((....))))).------------------- ( -38.70, z-score = -1.51, R) >consensus AGAUUUCACAGGC___AUUGGGCCCUGUUGAUUAACAUGGCAUGU_____UAAGUUCUUGGCCACAGAAAGCUUUUCGGAGUUAAUGGUUAAGAAAAGCUAUGGUAAUAGAGCCAG______GAAACUGGCAGCUUUCUAAAGCUC___________________ ........................((((((.....((((((.((......))..((((((((((.....(((((....)))))..))))))))))..)))))).)))))).(((((..........)))))(((((....))))).................... (-23.58 = -24.78 + 1.20)

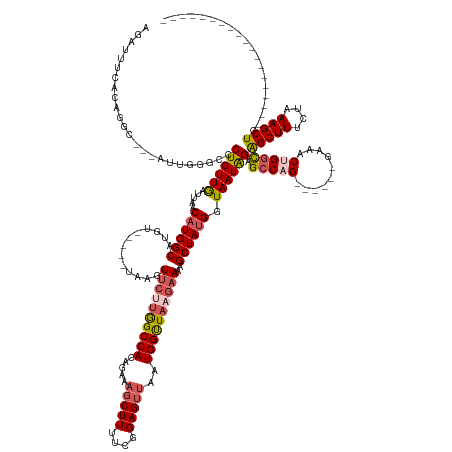

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:31:08 2011