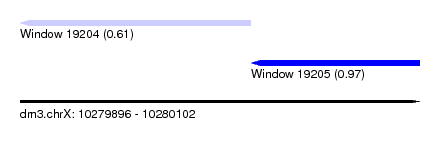
| Sequence ID | dm3.chrX |
|---|---|
| Location | 10,279,896 – 10,280,102 |
| Length | 206 |
| Max. P | 0.971530 |

| Location | 10,279,896 – 10,280,015 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.16 |
| Shannon entropy | 0.28393 |
| G+C content | 0.53786 |
| Mean single sequence MFE | -39.68 |
| Consensus MFE | -28.20 |
| Energy contribution | -28.00 |
| Covariance contribution | -0.20 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.71 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.608587 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
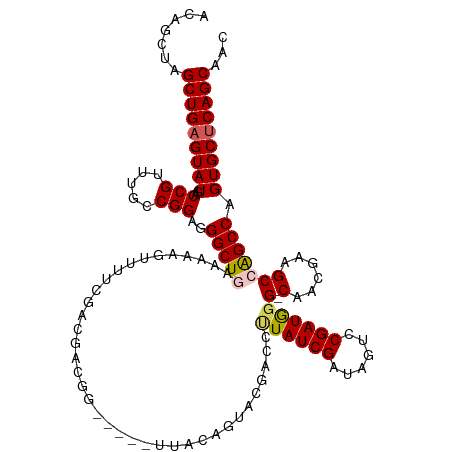
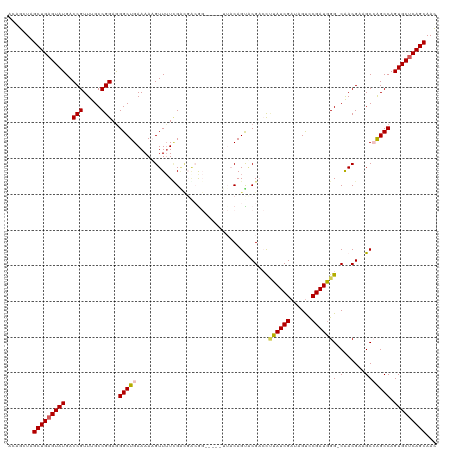
>dm3.chrX 10279896 119 - 22422827 ACAGCUAGCUGAGUAUGACCGUUUGCCGGAAGGCUGAAAAAGUUUUUGGUGGCGGCGGCUUUACAGUUCGACUUUAUCGAUAGUCCGAUGGG-CAACGAAGCCAGCCAGUGCUCAGCAAC .......(((((((((........(((((((((((.....)))))))..))))(((((((((.....((((.....))))..((((...)))-)...)))))).))).)))))))))... ( -44.20, z-score = -2.25, R) >droEre2.scaffold_4690 13892596 108 + 18748788 ACAGCUAGCUGAGUAUGACCGUUUGCCGGAGGGCUGAAAAAGUUAUUGGCGAGG---------CAGCGCG--CGUAUCGAUAGUUCGAUGGG-CAACGAAGCAGGCCAGUGCGCAGCAAC .(((((..(((.(((........))))))..))))).....(((((((((...(---------(..((.(--(.((((((....)))))).)-)..))..))..))))))).))...... ( -37.60, z-score = -0.79, R) >droYak2.chrX 18854252 109 - 21770863 ACAGCUAGCUGAGUAUGACCGUUUGCCGGAGGGCUGAAAAAGUUUUGGCAGUGG---------CAGCGCG--CCUAUCGAUAGCUCGAUAGGCCGACGAAGCAAGCCAGUGCUCAGCAAC .......(((((((((....((((((((..(.((((..(..((....))..)..---------)))).)(--((((((((....)))))))))...))..))))))..)))))))))... ( -45.00, z-score = -2.67, R) >droSec1.super_15 977452 114 - 1954846 ACAGCUAGCUGAGUAUGACCGUUUGCCGGAGGGCUGAAAAUGUUUUCGACUACAG-----UUACAGUACGACCUUAUCGAUAGUCCGAUGGG-CAACGCCGCCAGCCAGUGCUCAGCAAU .......(((((((((..(((.....)))..(((((....((((.((((((....-----....))).))).(((((((......)))))))-.))))....))))).)))))))))... ( -35.80, z-score = -1.29, R) >droSim1.chrX 8004318 114 - 17042790 ACAGCUAGCUGAGUAUGACCGUUUGCCGGAGGGCUGAAAAUGUUUUCGACUACAG-----UUACAGUACGACCUUAUCGAUAGUCCGAUGGG-CAACGCCGCCAGCCAGUGCUCAGCAAC .......(((((((((..(((.....)))..(((((....((((.((((((....-----....))).))).(((((((......)))))))-.))))....))))).)))))))))... ( -35.80, z-score = -1.25, R) >consensus ACAGCUAGCUGAGUAUGACCGUUUGCCGGAGGGCUGAAAAAGUUUUCGACGACGG_____UUACAGUACGACCUUAUCGAUAGUCCGAUGGG_CAACGAAGCCAGCCAGUGCUCAGCAAC .......(((((((((..(((.....)))..((((((((....))))..................((.((..(((((((......)))))))....))..)).)))).)))))))))... (-28.20 = -28.00 + -0.20)

| Location | 10,280,015 – 10,280,102 |
|---|---|
| Length | 87 |
| Sequences | 5 |
| Columns | 87 |
| Reading direction | reverse |
| Mean pairwise identity | 97.24 |
| Shannon entropy | 0.04721 |
| G+C content | 0.45496 |
| Mean single sequence MFE | -24.31 |
| Consensus MFE | -24.30 |
| Energy contribution | -24.14 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.93 |
| Structure conservation index | 1.00 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.85 |
| SVM RNA-class probability | 0.971530 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
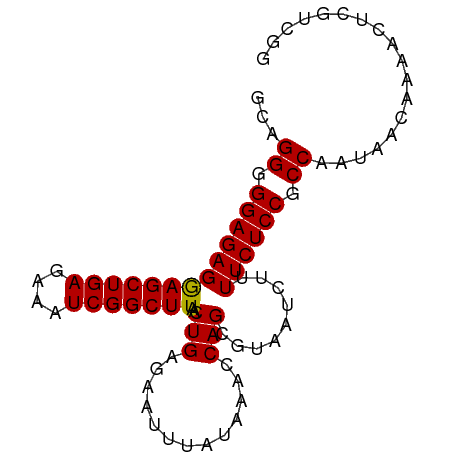
>dm3.chrX 10280015 87 - 22422827 GCAGGGGGAGAGGAGCUGAGAAAUCGGCUUAACUGAGAAUUUAUAAACCAGCGUAAUCUUUUCUCCGCCAAUAACAAAACUCGUCGG ...((.(((((((((((((....)))))))....((((..((((........)))))))))))))).)).................. ( -24.50, z-score = -1.90, R) >droEre2.scaffold_4690 13892704 86 + 18748788 GAAGGGGGAGAGGAGCUGAGAAAUCGGCUUAACUGAGAACUUAUAGACCAGCGUAAU-UUUUCUCCGCCAAUAACAAAACUCGUCGG ...((.(((((((((((((....)))))))..(((.............)))......-..)))))).)).................. ( -24.22, z-score = -1.72, R) >droYak2.chrX 18854361 86 - 21770863 GAAGGGGGAGAGGAGCUGAGAAAUCGGCUUAACUGAGAAUUUAUAAACCAGCGUAAU-UUUUCUCCGCCAAUAACAAAACUCGUCGG ...((.(((((((((((((....)))))))..(((.............)))......-..)))))).)).................. ( -24.22, z-score = -2.11, R) >droSec1.super_15 977566 87 - 1954846 GCAGGGGGAGAGAAGCUGAGAAAUCGGCUUAACUGAGAAUUUAUAAACCAGCGUAAUCUUUUCUCCGCCAAUAACAAAACUCGUCGG ...((.(((((((((((((....)))))))....((((..((((........)))))))))))))).)).................. ( -24.10, z-score = -2.02, R) >droSim1.chrX 8004432 87 - 17042790 GCAGGGGGAGAGGAGCUGAGAAAUCGGCUUAACUGAGAAUUUAUAAACCAGCGUAAUCUUUUCUCCGCCAAUAACAAAACUCGUCGG ...((.(((((((((((((....)))))))....((((..((((........)))))))))))))).)).................. ( -24.50, z-score = -1.90, R) >consensus GCAGGGGGAGAGGAGCUGAGAAAUCGGCUUAACUGAGAAUUUAUAAACCAGCGUAAUCUUUUCUCCGCCAAUAACAAAACUCGUCGG ...((.(((((((((((((....)))))))..(((.............))).........)))))).)).................. (-24.30 = -24.14 + -0.16)

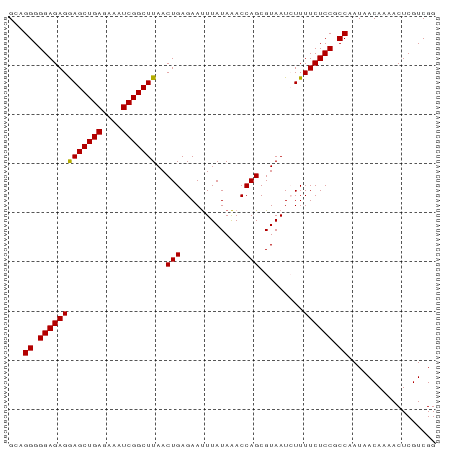
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:31:03 2011