| Sequence ID | dm3.chrX |
|---|---|
| Location | 9,965,663 – 9,965,763 |
| Length | 100 |
| Max. P | 0.961747 |

| Location | 9,965,663 – 9,965,763 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 66.71 |
| Shannon entropy | 0.62663 |
| G+C content | 0.60792 |
| Mean single sequence MFE | -27.47 |
| Consensus MFE | -13.26 |
| Energy contribution | -14.98 |
| Covariance contribution | 1.72 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.48 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.70 |
| SVM RNA-class probability | 0.961747 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
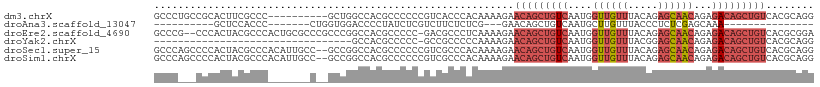
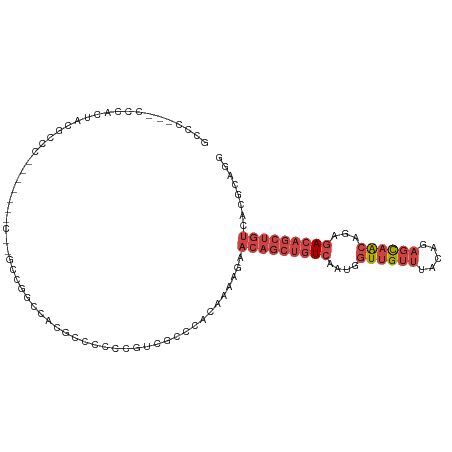
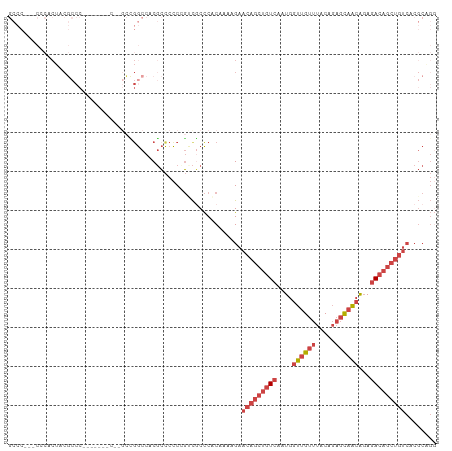
>dm3.chrX 9965663 100 + 22422827 GCCCUGCCGCACUUCGCCC----------GCUGGCCACGCCCCCCGUCACCCACAAAAGAACAGCUGUCAAUGGUUGUUUACAGAGCAACAGAGACAGCUGUCACGCAGG ..(((((.((.....(((.----------...)))...))..................((.((((((((....(((((((...)))))))...))))))))))..))))) ( -31.70, z-score = -1.81, R) >droAna3.scaffold_13047 1804450 75 - 1816235 ----------GCUCCACCC-------CUGGUGGACCCCUAUCUCGUCUUCUCUCG---GAACAGCUGUCAAUGCUUGUUUACCCUCUCGAGCAAA--------------- ----------..((((((.-------..)))))).......((((..........---(((((((.......)).))))).......))))....--------------- ( -15.93, z-score = -1.11, R) >droEre2.scaffold_4690 13606669 107 - 18748788 GCCCG--CCCACUACGCCCACUGCGCCCGCCCGGCCACGCCCCC-GACGCCCUCAAAAGAACAGCUGUCAAUGGUUGUUUACAGAGCAACAGAGACAGCUGUCACGCGGA ..(((--(..............(((..((...(((...)))..)-).)))........((.((((((((....(((((((...)))))))...))))))))))..)))). ( -33.90, z-score = -1.82, R) >droYak2.chrX 18555099 76 + 21770863 ---------------------------------GCCACGCCCCC-GCCGCCCCCAAAAGAACAGCUGUCAAUGGUUGUUUACGGAGCAACAGAGACAGCUGUCACGCAGG ---------------------------------((...((....-))...........((.((((((((....(((((((...)))))))...))))))))))..))... ( -22.30, z-score = -1.33, R) >droSec1.super_15 681696 108 + 1954846 GCCCAGCCCCACUACGCCCACAUUGCC--GCCGGCCACGCCCCCCGUCGCCCACAAAAGAACAGCUGUCAAUGGUUGUUUACAGAGCAACAGAGACAGCUGUCACGCAGG ......................((((.--...(((.(((.....))).))).......((.((((((((....(((((((...)))))))...))))))))))..)))). ( -30.50, z-score = -1.72, R) >droSim1.chrX 7933159 108 + 17042790 GCCCAGCCCCACUACGCCCACAUUGCC--GCCGGCCACGCCCCCCGUCGCCCACAAAAGAACAGCUGUCAAUGGUUGUUUACAGAGCAACAGAGACAGCUGUCACGCAGG ......................((((.--...(((.(((.....))).))).......((.((((((((....(((((((...)))))))...))))))))))..)))). ( -30.50, z-score = -1.72, R) >consensus GCCC___CCCACUACGCCC_______C__GCCGGCCACGCCCCCCGUCGCCCACAAAAGAACAGCUGUCAAUGGUUGUUUACAGAGCAACAGAGACAGCUGUCACGCAGG ............................................................(((((((((....((((((.....))))))...)))))))))........ (-13.26 = -14.98 + 1.72)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:30:07 2011