
| Sequence ID | dm3.chrX |
|---|---|
| Location | 9,899,684 – 9,899,774 |
| Length | 90 |
| Max. P | 0.657639 |

| Location | 9,899,684 – 9,899,774 |
|---|---|
| Length | 90 |
| Sequences | 13 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 65.03 |
| Shannon entropy | 0.72958 |
| G+C content | 0.48828 |
| Mean single sequence MFE | -26.30 |
| Consensus MFE | -11.85 |
| Energy contribution | -12.33 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.42 |
| Mean z-score | -0.66 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.657639 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 9899684 90 - 22422827 ----ACCUUAGAUCAGCGCCGUUAUCAGCUUGGCCUUGAAGUUCACGAACUUCUCGCUGGUGGCCGCAACGCGCUGCAAACGGGUAGGUAACAU--------------------- ----((((.....((((((.(((((((((........((((((....))))))..)))))))))......)))))).....)))).........--------------------- ( -27.10, z-score = -0.14, R) >droSec1.super_130 51147 90 - 56455 ----ACCUUAGAUCAGCGCCGUUAUCAGCUUGGCCUUGAAGUUCACGAACUUCUCGCUGGUGGCCGCAACGCGCUGCAAACGGGAAGGUAACAU--------------------- ----(((((.(..((((((.(((((((((........((((((....))))))..)))))))))......))))))....)...))))).....--------------------- ( -27.00, z-score = -0.19, R) >droYak2.chrX 18491305 88 - 21770863 ----ACCUUAGAUCAGCGCCGUUAUCAGCUUGGCCUUGAAGUUCACGAACUUCUCGCUGGUGGCCGCAACGCGCUGCAAACGA--AGGCGAUUG--------------------- ----.((((....((((((.(((((((((........((((((....))))))..)))))))))......))))))......)--)))......--------------------- ( -26.60, z-score = -0.09, R) >droEre2.scaffold_4690 13539705 90 + 18748788 ----ACCUUAGAUCAGCGCCGUUAUCAGCUUGGCCUUGAAGUUCACGAACUUCUCGCUGGUGGCCGCCACGCGCUGCAAGAGGGAAGCCAAUCA--------------------- ----.((((....((((((.(((((((((........((((((....))))))..)))))))))......))))))....))))..........--------------------- ( -28.20, z-score = -0.05, R) >droAna3.scaffold_13047 266502 115 - 1816235 ACCUUUCUUAAAUCAGCGCCGUUAUGAGCUUGGCCUUGAAGUUCACGAACUUCUCGCUGGUGGCGGCCACGCGCUGAAAGAGAGAGGAGGAACAGGUUAGCACAAGGGGCAACAU .((((((((...(((((((.((.....)).(((((..((((((....))))))..((.....))))))).)))))))..))))))))........(((.((.......))))).. ( -40.40, z-score = -1.49, R) >dp4.chrXL_group3a 1132725 106 - 2690836 ----GCCUUAGAUCAGCGCCGUUAUCAGCUUGGCCUUGAAGUUGACGAACUUCUCGCUGGUGGCGGCCACGCGCUGCAAAUAGAAGGAAAAGUAUACGAUACGAUUACAA----- ----.((((..((((((((((((((((((........((((((....))))))..)))))))))))).....))))...))..))))....((((...))))........----- ( -30.60, z-score = -0.52, R) >droPer1.super_15 421896 106 + 2181545 ----UCCUUAGAUCAGCGCCGUUAUCAGCUUGGCCUUGAAGUUGACGAACUUCUCGCUGGUGGCGGCCACGCGCUGCAAUUAGAAGGAAAAGUAUACGAUACGAUCACAU----- ----(((((....((((((((((((((((........((((((....))))))..)))))))))))).....)))).......)))))...((((...))))........----- ( -31.60, z-score = -0.94, R) >droWil1.scaffold_181096 9439675 90 + 12416693 ----AGCCUAAAUCAACGCCGUUAUCAAUUUGGCCUUAAAGUUGACAAACUUCUCGCUGGUUGCGGCAACACGCUAAAACAAAAAAGACAAAUU--------------------- ----.(((..(((((.(((((.........))))....(((((....)))))...).)))))..)))...........................--------------------- ( -14.50, z-score = 0.21, R) >droVir3.scaffold_12970 3053760 84 + 11907090 ------AUUAUAUCAGCGCCGUUAUCAAUUUGGCCUUGAAGUUCACAAACUUUUCGCUCGUGGCGGCAACACGCUAUGGA----UAUUGACAGU--------------------- ------...((((((((((((.........))))...((((((....))))))..)))(((((((......)))))))))----))).......--------------------- ( -21.40, z-score = -1.25, R) >droMoj3.scaffold_6473 16652738 84 - 16943266 ------CUUAUAUCAGGGCCGUUAUCAAUUUGGCCUUGAAGUUCACGAACUUCUCGCUCGUGGCGGCGACGCGCUAAGAAAUGAGAUAAG------------------------- ------(((((.(((((((((.........)))))))))(((.(((((.(.....).)))))(((....)))))).....))))).....------------------------- ( -27.80, z-score = -2.37, R) >droGri2.scaffold_15203 430676 91 + 11997470 ------CUUAGAUCAGCGCCGUUAUCAAUUUUGCCUUGAAGUUGACAAACUUCUCGCUGGUGGCGGCAACGCGCUAUAGAAUUGCAUUGGGUAUAAU------------------ ------(((((....((((((((((((.....((...((((((....))))))..))))))))))))...))((.........)).)))))......------------------ ( -23.60, z-score = -0.40, R) >anoGam1.chrX 12536443 111 - 22145176 ----AUCUUAGAUGAGUGACGUAAUUAGGCGUGCUUUAAAGUUAACCACUCGUUCACUGGUUUGUGCGAUUCGCUGUGAAAGAAAAAGAAAGGGGAGGCACAAACAUGGGAAUGU ----.(((((((((((((((((......))))(((....)))....)))))))).....((((((((..(((.((...............)).))).)))))))).))))).... ( -29.16, z-score = -1.46, R) >triCas2.ChLG4 7632020 87 - 13894384 ------CUUAGAUGAGAGAUGUAACAAGACGAGCCUUGAAAUUGACUAAUCUUUCGCUAGUUUGGGCCAAACGCUGCAACAAAAAAACGAAUC---------------------- ------..(((.((((((((.(((((((......))))...)))....)))))))))))(((..(((.....)))..))).............---------------------- ( -13.90, z-score = 0.06, R) >consensus ____ACCUUAGAUCAGCGCCGUUAUCAGCUUGGCCUUGAAGUUCACGAACUUCUCGCUGGUGGCGGCAACGCGCUGCAAAAGGAAAGAAAAUUU_____________________ ..............(((((.((..(((..(..((...((((((....))))))..))..))))..))...)))))........................................ (-11.85 = -12.33 + 0.48)


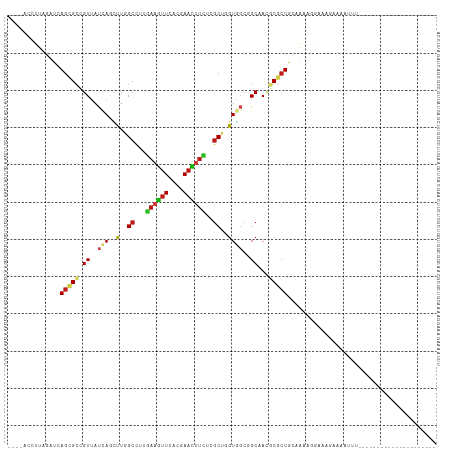
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:30:01 2011