
| Sequence ID | dm3.chrX |
|---|---|
| Location | 9,840,070 – 9,840,172 |
| Length | 102 |
| Max. P | 0.741439 |

| Location | 9,840,070 – 9,840,172 |
|---|---|
| Length | 102 |
| Sequences | 7 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 56.43 |
| Shannon entropy | 0.87180 |
| G+C content | 0.50355 |
| Mean single sequence MFE | -30.92 |
| Consensus MFE | -10.13 |
| Energy contribution | -10.23 |
| Covariance contribution | 0.10 |
| Combinations/Pair | 1.82 |
| Mean z-score | -1.05 |
| Structure conservation index | 0.33 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.741439 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 9840070 102 + 22422827 -------AGUUUCUCGGCUCUUUUUUCCGGGCCGAGAACAAUGUCGGCAUAGUUGGUCGAUGGGCGGAGCGAGCAGAAAUUGCCCGGGGGCAAGAAGGCCAACUAAUGG -------.(((.((((((((........)))))))))))........(((((((((((.....((.......)).....(((((....)))))...)))))))).))). ( -39.70, z-score = -1.86, R) >droWil1.scaffold_181096 9355043 97 - 12416693 ----AAUAUCUUUUUGGU--UUUCUUAAGAGUCUUGGUUUCCCUCGAAG----UCCACAAAUUUUUUUUCGGUUUAAAGAUGAUCCUGAUUAUUAUAAUCUAAAUAC-- ----..(((((((.....--......((((((..(((((((....))))----.)))...)))))).........))))))).....(((((...))))).......-- ( -9.85, z-score = 1.62, R) >droSim1.chrX 7838802 99 + 17042790 -------AGUUUCUCGGUUCUUUUUUCCGGGCCGAGAACAAUGUCGGCA--GUUGGUCGAUGGGCGGAGCCAGUAGAAAUUGGCCGGGG-CAAGAAGGCCAACUAAUGG -------.(((.((((((((........))))))))))).....((..(--(((((((.....((...((((((....))))))....)-).....))))))))..)). ( -36.10, z-score = -1.59, R) >droSec1.super_15 604792 99 + 1954846 -------AGUUUCUAGGUUCUUUUUUCCGGGCCGAGAACAAUGUCGGCA--GUUGGUCGAUGGGCGGAGCCAGUAGAAAUUGGCCGGGG-CAAGAAGGCCAACUAAUGG -------........(((..((((.(((.((((((.......((((((.--....)))))).(((...)))........)))))).)))-.))))..)))......... ( -33.00, z-score = -0.95, R) >droYak2.chrX 18433019 89 + 21770863 AGUUUCUCAUUUCUCGGUUCUUUUUUCCGUGCCGAGAACAAUGUCAGUA-------------------GCCAGUGGAAAUUGGCCGAGA-GAGGAAGGCCAACUAAUGG ..((((((..((((((((.(........).))))))))...........-------------------((((((....)))))).))))-))((....))......... ( -27.60, z-score = -1.75, R) >droEre2.scaffold_4690 13478492 99 - 18748788 -------AGUUUCUCGGUUCUGUUUUCCGGGCCGAGAACAAUGUCAGUA--GUUGGUCGAUGGGCGGAGCCAUGGGAAAUUAGCCGGGG-AAAGAAGACCAAAUAAUGG -------..((((((((((..(((((((.((((((..((.......)).--.)))))).((((......))))))))))).))))))))-))......(((.....))) ( -31.30, z-score = -1.32, R) >droAna3.scaffold_13047 218126 92 + 1816235 -------AGCUUCCUGCUCCUCCCUCCCGCGUGGUGG-----GUCCGAAACACCGUACGGGGAAGGUCUCCAGCCACAAGGUUGCAGGACGAGGAGUGUAGUGU----- -------.(((...((((((((..(((((..(((((.-----........)))))..)))))...((((.(((((....)))))..)))))))))))).)))..----- ( -38.90, z-score = -1.48, R) >consensus _______AGUUUCUCGGUUCUUUUUUCCGGGCCGAGAACAAUGUCGGCA__GUUGGUCGAUGGGCGGAGCCAGUAGAAAUUGGCCGGGG_CAAGAAGGCCAACUAAUGG ...............(((..(((((((((((((((.(....).)))))........(((.....)))................))))))..))))..)))......... (-10.13 = -10.23 + 0.10)
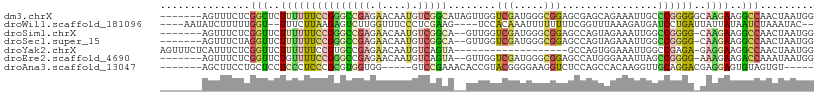

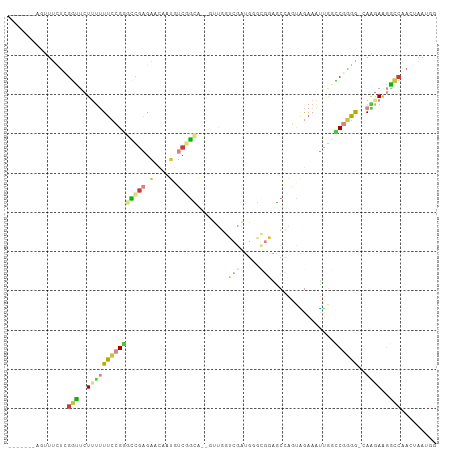
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:29:51 2011