
| Sequence ID | dm3.chrX |
|---|---|
| Location | 9,731,199 – 9,731,290 |
| Length | 91 |
| Max. P | 0.545573 |

| Location | 9,731,199 – 9,731,290 |
|---|---|
| Length | 91 |
| Sequences | 11 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 84.86 |
| Shannon entropy | 0.32613 |
| G+C content | 0.38818 |
| Mean single sequence MFE | -19.54 |
| Consensus MFE | -15.40 |
| Energy contribution | -15.78 |
| Covariance contribution | 0.39 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.10 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.545573 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 9731199 91 + 22422827 GCCCGCUCAAAUAUUAACACAGAUAAACGAGCGGAAGUGUAUUUGAUCAGCAUUUAAUGCAG----CAAAUGUCGAUUUGUCGCUGA--CAAAAAAC-- ..((((((...((((......))))...)))))).((((.....((((.((((((.......----.)))))).))))...))))..--........-- ( -20.90, z-score = -1.57, R) >droSim1.chrX 7758843 91 + 17042790 GCCCGCUCAAAUAUUUACACAGAUAAACGAGCGGAAGUGUAUUUGAUCAGCAUUUAAUGCAG----CAAAUGUCGAUUUGUCGCUGA--CAAAAAAC-- ..((((((...(((((....)))))...)))))).((((.....((((.((((((.......----.)))))).))))...))))..--........-- ( -21.10, z-score = -1.52, R) >droSec1.super_15 486762 91 + 1954846 GCCCGCUCAAAUAUUUACACAGAUAAACGAGCGGAAGUGUAUUUGAUCAGCAUUUAAUGCAG----CAAAUGUCGAUUUGUCGCUGA--CAAAAAAC-- ..((((((...(((((....)))))...)))))).((((.....((((.((((((.......----.)))))).))))...))))..--........-- ( -21.10, z-score = -1.52, R) >droYak2.chrX 18323922 91 + 21770863 GCCCGCUCAAAUAUUUACACAGAUAAACGAGCGGAAGUGUAUUUGAUCAGCAUUUAAUGCAG----CAAAUGUCGAUUUGUCGCUGA--CAAAAAAA-- ..((((((...(((((....)))))...)))))).((((.....((((.((((((.......----.)))))).))))...))))..--........-- ( -21.10, z-score = -1.46, R) >droEre2.scaffold_4690 13360578 91 - 18748788 GCCCGCUCAAAUAUUUACACAGAUAAACGAGCGGAAGUGUAUUUGAUCAGCAUUUAAUGCAG----CGAAUGUCGAUUUGUCGCUGA--CAAAAAAC-- ..((((((...(((((....)))))...))))))((((((.........))))))..(((((----(((...........)))))).--))......-- ( -21.90, z-score = -1.56, R) >droAna3.scaffold_13047 1467001 91 - 1816235 GUCCGUUCAAAUAUUUACCCAGAUAAACGAGCGGAAGUGUAUUUGAUCAGCAUUUAAUGCAG----CAAAUGUCGAUUUGUCGCUGA--CAAAAAAC-- .(((((((...(((((....)))))...)))))))((((.....((((.((((((.......----.)))))).))))...))))..--........-- ( -20.20, z-score = -1.54, R) >dp4.chrXL_group3a 959289 91 + 2690836 GCCUGCUCAAAUAUUUACACAGAUAAACGAGCGGAAGUGUAUUUGAUCAGCAUUUAACGUAG----CAAAUGUCGAUUUGUCGCUGA--CAAAAAAA-- ..((((((...(((((....)))))...)))))).((((.....((((.((((((.......----.)))))).))))...))))..--........-- ( -18.90, z-score = -0.71, R) >droPer1.super_15 243367 91 - 2181545 GCCUGCUCAAAUAUUUACACAGAUAAACGAGCGGAAGUGUAUUUGAUCAGCAUUUAACGUAG----CAAAUGUCGAUUUGUCGCUGA--CAAAAAAA-- ..((((((...(((((....)))))...)))))).((((.....((((.((((((.......----.)))))).))))...))))..--........-- ( -18.90, z-score = -0.71, R) >droWil1.scaffold_181096 9220815 95 - 12416693 GCCUGCUCAAAUAUUUAUACAGAUAAACGAGCGGAAGUGUAUUUGAUCAGCAUUUAACGUAG----CAAAUGUCGAUUUGUCGCAAAGUCGCAAAAAAC ((((((((...(((((....)))))...))))))..(((.....((((.((((((.......----.)))))).))))...)))......))....... ( -19.80, z-score = -0.82, R) >droMoj3.scaffold_6473 16373874 80 + 16943266 GUCUACUUAAAUAUUUACGCCGAUAAACGAGCG---GCUUAUUUGAUCAAGGUUAAAGUCAA----CAUUUGA-AGUUCACUGCCAAA----------- ......(((((((....(((((.....)).)))---...)))))))....(((...(((.((----(......-.))).))))))...----------- ( -13.10, z-score = -0.40, R) >droGri2.scaffold_15203 233505 86 - 11997470 GACUGCUCAAAUAUUUACACAGAUAAACGAGCGGAAGUGUAUUUGAUCAGCAGUUAACGUACUUGCCUAAUGCCGAUUUGUCGCUG------------- ..((((((...(((((....)))))...)))))).((((.....((((.(((.....((....)).....))).))))...)))).------------- ( -17.90, z-score = -0.32, R) >consensus GCCCGCUCAAAUAUUUACACAGAUAAACGAGCGGAAGUGUAUUUGAUCAGCAUUUAAUGCAG____CAAAUGUCGAUUUGUCGCUGA__CAAAAAAC__ ..((((((...((((......))))...)))))).((((.....((((.((((((............)))))).))))...)))).............. (-15.40 = -15.78 + 0.39)

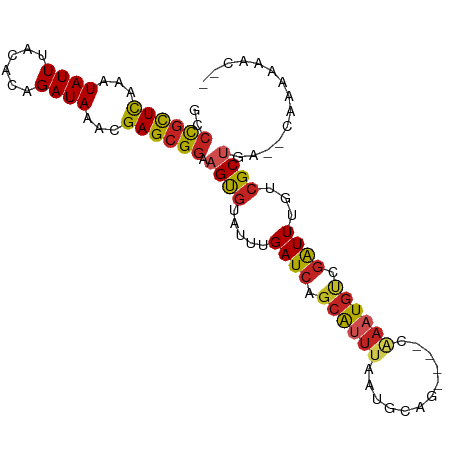

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:29:35 2011