
| Sequence ID | dm3.chrX |
|---|---|
| Location | 9,714,350 – 9,714,411 |
| Length | 61 |
| Max. P | 0.525521 |

| Location | 9,714,350 – 9,714,411 |
|---|---|
| Length | 61 |
| Sequences | 9 |
| Columns | 64 |
| Reading direction | reverse |
| Mean pairwise identity | 72.66 |
| Shannon entropy | 0.55470 |
| G+C content | 0.39352 |
| Mean single sequence MFE | -12.69 |
| Consensus MFE | -6.07 |
| Energy contribution | -5.99 |
| Covariance contribution | -0.09 |
| Combinations/Pair | 1.40 |
| Mean z-score | -1.11 |
| Structure conservation index | 0.48 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.525521 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
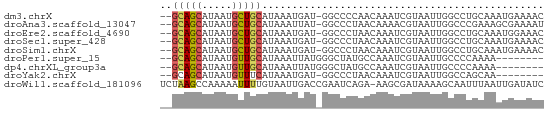
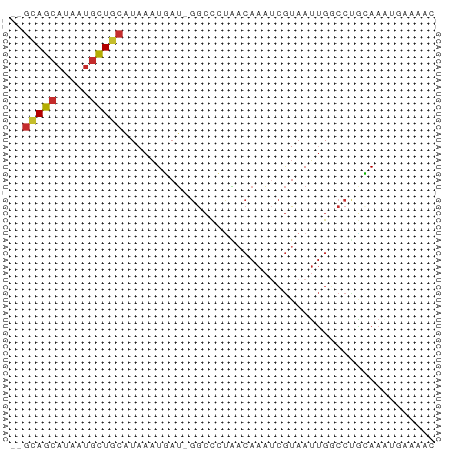
>dm3.chrX 9714350 61 - 22422827 --GCAGCAUAAUGCUGCAUAAAUGAU-GGCCCCAACAAAUCGUAAUUGGCCUGCAAAUGAAAAC --(((((.....))))).....((..-((((...((.....))....))))..))......... ( -14.90, z-score = -1.59, R) >droAna3.scaffold_13047 1450424 61 + 1816235 --GCAGCAUAAUGCUGCAUAAAUUAU-GGCCCUAACAAAACGUAAUUGGCCCGAAAGCGAAAAU --(((((.....))))).........-((((...((.....))....))))(....)....... ( -14.70, z-score = -1.85, R) >droEre2.scaffold_4690 13343844 61 + 18748788 --GCAGCAUAAUGCUGCAUAAAUGAU-GGCCCUAACAAAUCGUAAUUGGCCUGCAAAUGGAAAC --(((((.....))))).....((..-((((...((.....))....))))..))......... ( -14.90, z-score = -1.24, R) >droSec1.super_428 5380 61 - 11857 --GCAGCAUAAUGCUGCAUAAAUGAU-GGCCCUAACAAAUCGUAAUUGGCCUGCAAAUGAAAAC --(((((.....))))).....((..-((((...((.....))....))))..))......... ( -14.90, z-score = -1.83, R) >droSim1.chrX 7745510 61 - 17042790 --GCAGCAUAAUGCUGCAUAAAUGAU-GGCCCUAACAAAUCGUAAUUGGCCUGCAAAUGAAAAC --(((((.....))))).....((..-((((...((.....))....))))..))......... ( -14.90, z-score = -1.83, R) >droPer1.super_15 227089 54 + 2181545 --GCAGCAUAAUGUUGCAUAAAUUAUGGGCUAUGCCAAAUCGUAAUUGCCCCAAAA-------- --(((((.....))))).........((((..(((......)))...)))).....-------- ( -14.00, z-score = -1.46, R) >dp4.chrXL_group3a 944407 54 - 2690836 --GCAGCAUAAUGUUGCAUAAAUUAUGGGCUAUGCCAAAUCGUAAUUGCCCCAAAA-------- --(((((.....))))).........((((..(((......)))...)))).....-------- ( -14.00, z-score = -1.46, R) >droYak2.chrX 18307009 53 - 21770863 --GCAGCAUAAUGUUUCAUAAAUGAU-GGCCCUAACAAAUCGUAAUUGGCCAGCAA-------- --...((........(((....)))(-((((...((.....))....)))))))..-------- ( -9.50, z-score = -0.20, R) >droWil1.scaffold_181096 9201786 63 + 12416693 UCUAAGCCAAAAAUUUUGUAAUUGACCGAAUCAGA-AAGCGAUAAAAGCAAUUUAAUUGAUAUC .............(((((.............))))-)..((((.(((....))).))))..... ( -2.42, z-score = 1.42, R) >consensus __GCAGCAUAAUGCUGCAUAAAUGAU_GGCCCUAACAAAUCGUAAUUGGCCUGCAAAUGAAAAC ..(((((.....)))))............................................... ( -6.07 = -5.99 + -0.09)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:29:32 2011