
| Sequence ID | dm3.chrX |
|---|---|
| Location | 9,691,917 – 9,692,035 |
| Length | 118 |
| Max. P | 0.902301 |

| Location | 9,691,917 – 9,692,035 |
|---|---|
| Length | 118 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.74 |
| Shannon entropy | 0.35060 |
| G+C content | 0.47978 |
| Mean single sequence MFE | -34.24 |
| Consensus MFE | -20.45 |
| Energy contribution | -21.70 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.16 |
| SVM RNA-class probability | 0.902301 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 9691917 118 - 22422827 AGAACUUCGCUUUCCGGCCAAGUUGAGCCAACUAGUUUGUUGGCUUCGCUUCAACUUACUUUU--GGCCAUUUUACGCUGGAUUUAGCACUUGGCCACAGAUCGUCCAGAUGAUGCGCAA .......(((.....((((((((.((((((((......)))))))).(((..........((.--.((........))..))...)))))))))))....(((((....))))))))... ( -36.92, z-score = -2.34, R) >droAna3.scaffold_13047 1438385 95 + 1816235 AAAAGUUCGC-CCCCGGCCAAGUUGAUCAAACUAAUUUGUUGGCUUC---CCAACUUUC------GGCUGCCGGGGAGGGAUCUUGCCAC---ACAAUAAAUCCGCAA------------ ........((-(((((((..((((((............(((((....---)))))..))------)))))))))))..((((.(((....---.)))...))))))..------------ ( -29.24, z-score = -1.28, R) >droEre2.scaffold_4690 13329165 118 + 18748788 AGAACUUCGCUUUCCGGCCAAGUUGAGCCAACUAGUUUGUUGGCUUCGCCUCAACUUACUUUU--GGCCAUUUUACGCUGGAUUUACCACUUGGCCACAGAUCGUCCAGAUGAUGCACAA ........((.....((((((((.((((((((......))))))))..............((.--.((........))..))......))))))))....(((((....))))))).... ( -34.60, z-score = -2.52, R) >droYak2.chrX 18295036 120 - 21770863 AGAACUUCGCUUUCCGGCCAAGUUGAGCCAACUAGUUUGUUGGCUUCGCUUCAACUUACUUUUUGGGCCAUUUUACGCUGGAUUUAGCACUUGGCCACAGAUCGCCCAGAUGAUGUACUA ((.((.(((((.((((((.((((.((((((((......)))))))).)))).......((....))..........))))))...))).((.(((........))).))..)).)).)). ( -35.00, z-score = -1.95, R) >droSec1.super_15 455497 118 - 1954846 AGAACUUCGCUUUCCGGCCAAGUUGAGCCAACUAAUUCGUUGGCUUCGCUUCAACUUACUUUU--GGCCAUUUUACGCUGGACUUACCACUUGGCCACAGAUCGUCCAGAUGAUGCGCAA .......(((.....((((((((.((((((((......))))))))..............((.--.((........))..))......))))))))....(((((....))))))))... ( -34.80, z-score = -2.58, R) >droSim1.chrX 7730178 118 - 17042790 AGAACUUCGCUUUCCGGCCAAGUUGAGCCAACUAAUUCGUUGGCUUCGCUUCAACUUACUUUU--GGCCAUUUUACGCUGGAUUUACCACUUGGCCACAGAUCGUCCAGAUGAUGCGCAA .......(((.....((((((((.((((((((......))))))))..............((.--.((........))..))......))))))))....(((((....))))))))... ( -34.90, z-score = -2.62, R) >consensus AGAACUUCGCUUUCCGGCCAAGUUGAGCCAACUAAUUUGUUGGCUUCGCUUCAACUUACUUUU__GGCCAUUUUACGCUGGAUUUACCACUUGGCCACAGAUCGUCCAGAUGAUGCGCAA ........((.....(((((((((((((((((......))))))......)))))))........))))........((((((....................)))))).....)).... (-20.45 = -21.70 + 1.25)

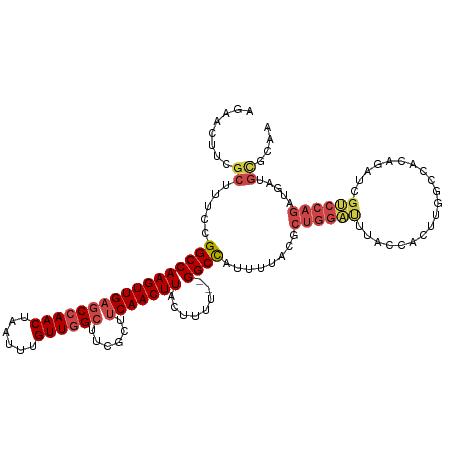
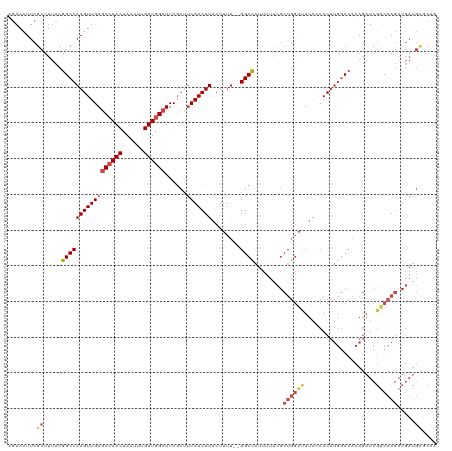
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:29:29 2011