
| Sequence ID | dm3.chrX |
|---|---|
| Location | 9,635,766 – 9,635,866 |
| Length | 100 |
| Max. P | 0.581894 |

| Location | 9,635,766 – 9,635,866 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 78.38 |
| Shannon entropy | 0.39503 |
| G+C content | 0.40755 |
| Mean single sequence MFE | -23.75 |
| Consensus MFE | -13.58 |
| Energy contribution | -14.08 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.57 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.581894 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 9635766 100 - 22422827 ---------CUGAAAAUACUGUAGACGGGAGACAGAAGACAGAAGACCGGUAGCCAAUUGCUUUGGCAGUUAAGGUUGCACCAAUUCGUUUAAUAAUAAAUAAUGAAAA ---------.........((((.....(....).....)))).......((((((((((((....))))))..)))))).....((((((...........)))))).. ( -21.50, z-score = -1.76, R) >droSim1.chrX 7682548 99 - 17042790 ---------CUGAAAA-ACUGUAGACGGGAGACAGAAGACAGAAGACCGGUAGCCAAUUGCUUUGGCAGUUAAGGUUGCACCAAUUCGUUUAAUAAUAAAUAAUGAAAA ---------.......-.((((.....(....).....)))).......((((((((((((....))))))..)))))).....((((((...........)))))).. ( -21.50, z-score = -1.70, R) >droSec1.super_15 398054 99 - 1954846 ---------CUGAAAA-ACUGUAGACGGGAGACAGAAGACAGAAGACCGGUAGCCAAUUGCUUUGGCAGUUAAGGUUGCACCAAUUCGUUUAAUAAUAAAUAAUGAAAA ---------.......-.((((.....(....).....)))).......((((((((((((....))))))..)))))).....((((((...........)))))).. ( -21.50, z-score = -1.70, R) >droAna3.scaffold_13248 1463889 84 - 4840945 -------------------------GAAAAGAUGGCUGGAAGAAGACCGGUAGCCAAUUGCUUUGGCAGUUAAGGUUGCACCAAUCCGUUUAAUAAUAAAUAAUGAAAA -------------------------....((((((.(((..(.....).((((((((((((....))))))..)))))).)))..)))))).................. ( -18.60, z-score = -1.26, R) >droPer1.super_15 149482 108 + 2181545 GGGACUGGCGAAUGGCGAAUGGCAGUCGGAGAAGAGUGG-AGAAGACCGGUAGCCAAUUGCUUUGGCAGCUAAGGUUGCACCAAUUCGUUUAAUAAUAAAUAAUGAAAA ......((((((((((((.((((..((((..........-......))))..)))).)))))...(((((....)))))....)))))))................... ( -27.79, z-score = -1.09, R) >dp4.chrXL_group3a 864569 108 - 2690836 GGGACUGGCGAAUGGCGAAUGGCAGCCGGAGAAGAGUGG-AGAAGACCGGUAGCCAAUUGCUUUGGCAGCUAAGGUUGCACCAAUUCGUUUAAUAAUAAAUAAUGAAAA ......((((((((((((.((((.(((((..........-......))))).)))).)))))...(((((....)))))....)))))))................... ( -31.59, z-score = -2.06, R) >consensus _________CUGAAAA_ACUGUAGACGGGAGACAGAAGACAGAAGACCGGUAGCCAAUUGCUUUGGCAGUUAAGGUUGCACCAAUUCGUUUAAUAAUAAAUAAUGAAAA .........................................(((.....((((((((((((....))))))..)))))).....)))...................... (-13.58 = -14.08 + 0.50)
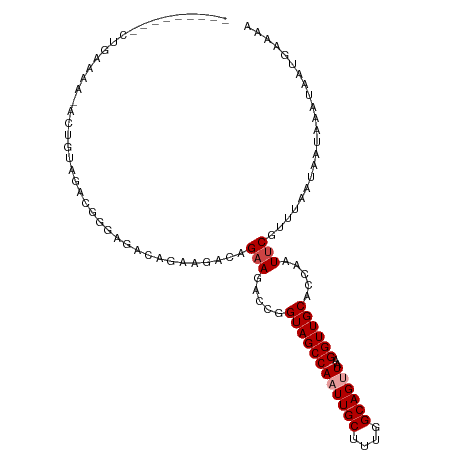


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:29:20 2011