
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 10,766,862 – 10,766,959 |
| Length | 97 |
| Max. P | 0.741521 |

| Location | 10,766,862 – 10,766,959 |
|---|---|
| Length | 97 |
| Sequences | 8 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 74.27 |
| Shannon entropy | 0.45803 |
| G+C content | 0.61641 |
| Mean single sequence MFE | -40.48 |
| Consensus MFE | -24.17 |
| Energy contribution | -22.50 |
| Covariance contribution | -1.67 |
| Combinations/Pair | 1.56 |
| Mean z-score | -1.35 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.741521 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 10766862 97 + 23011544 ---------------UCCGGCAGUCGUCGGCUGGCUGGUAUUGCUCACGUUGCUGGCCGUUGUGCUGGGUGCACUGGGCGCAUUUGAGGUGGCACUGGAGGUGGCCACCUCG ---------------.(((((....)))))..(((..(((.((....)).)))..)))..((((((.((....)).))))))...((((((((((.....)).)))))))). ( -45.20, z-score = -1.30, R) >droSim1.chr2L 10567958 97 + 22036055 ---------------UCCGGCAGACGUUGGCUGGCUGGGAUUGCUCACGUUGCUGGUCGUUGUGCUGGGUGCACUGGCGCUCAUUGAGGUGGCACUGGAGGUGGCCACCUCG ---------------.(((((((((((..((.(((((((....)))).)))))..).)))).))))))((((....)))).....((((((((((.....)).)))))))). ( -43.80, z-score = -1.39, R) >droSec1.super_3 6185355 97 + 7220098 ---------------UCCGGCAGACGUCGGCUGGCUGGGAUUGCUCACGUUGCUGGUCGUUGUGCUGGGUGCACUGGGAGCAUUUGGGGUGGCACUGGAGGUGGCCAUCUCG ---------------.(((((((((((((((.(((((((....)))).)))))))).)))).))))))((((.......))))..((((((((((.....)).)))))))). ( -40.00, z-score = -0.74, R) >droYak2.chr2L 7172052 97 + 22324452 ---------------UCCGGCAGACGUCGGCUGGCUGGGAUUGUUCACAUUGCUGGCCGUUGUGCUGGGUGCACUGGGCGCAUUUGAGGUGGCACUGGAGGUGGCCACUUCG ---------------.((((((.(...((((..((..(((....)).)...))..)))).).))))))((((.....))))....((((((((((.....)).)))))))). ( -44.30, z-score = -1.98, R) >droEre2.scaffold_4929 11976950 97 - 26641161 ---------------UCCGGCAGACGUCGGCUGGCUGGGAUUGUUCACAUUGCUGGCCGUUGUGCUGGGUGCACUAGGCGCAUUUGAGGUGGCACUGGAGGUGGCCACCUCG ---------------.((((((.(...((((..((..(((....)).)...))..)))).).))))))((((.....))))....((((((((((.....)).)))))))). ( -47.00, z-score = -2.49, R) >dp4.chr4_group4 2970216 85 + 6586962 ---------------UCCGGUGGAUGUGGGUUG------ACUGUUGGCAGUGCUGGC---UGUGCUGGAGGCGCUGGA---AUUGGAGCUGGCGCUGGAGGUUGCCACUUCU ---------------(((((((.(.((.(((..------((((....))))))).))---).)))))))(((((..(.---.......)..)))))((((((....)))))) ( -32.40, z-score = -0.93, R) >droPer1.super_10 1985240 85 + 3432795 ---------------UCCGGUGGAUGUGGGUUG------ACUGUUGGCAGUGCUGGC---UGUGCUGGAGGCGCUGGA---AUUAGAGCUGGCGCUGGAGGUUGCCACUUCU ---------------(((((((.(.((.(((..------((((....))))))).))---).)))))))(((((..(.---.......)..)))))((((((....)))))) ( -32.40, z-score = -0.95, R) >droWil1.scaffold_180703 301235 103 - 3946847 UGCUGUGGCUGGGCCACCGGCAGUUGUUACCUG------AUUAUUGGCAUUACUGGCUGUUGUGCUUGGAGUGCUGGA---AUUUGAGCUAGCACUGGAGGUGGCCACUUCA .((....))..(((((((..((((.((((.(((------(((.(..(((((...(((......)))...)))))..))---)))..)).))))))))..)))))))...... ( -38.70, z-score = -1.01, R) >consensus _______________UCCGGCAGACGUCGGCUGGCUGGGAUUGUUCACAUUGCUGGCCGUUGUGCUGGGUGCACUGGG_GCAUUUGAGGUGGCACUGGAGGUGGCCACCUCG ................(((((((.(((.(((...........))).)))))))))).....((((.....))))..........((((.((((((.....)).)))).)))) (-24.17 = -22.50 + -1.67)


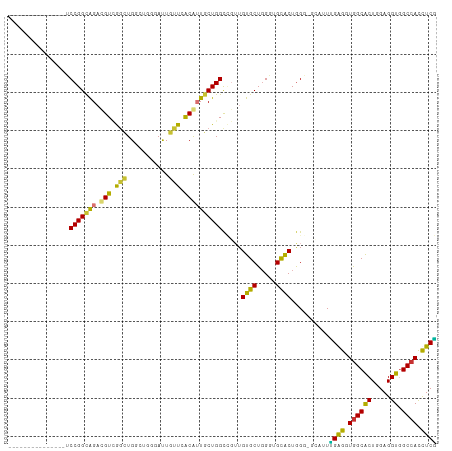
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:30:55 2011