
| Sequence ID | dm3.chrX |
|---|---|
| Location | 9,576,226 – 9,576,320 |
| Length | 94 |
| Max. P | 0.781585 |

| Location | 9,576,226 – 9,576,320 |
|---|---|
| Length | 94 |
| Sequences | 9 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 66.85 |
| Shannon entropy | 0.66258 |
| G+C content | 0.66110 |
| Mean single sequence MFE | -38.82 |
| Consensus MFE | -19.09 |
| Energy contribution | -18.49 |
| Covariance contribution | -0.60 |
| Combinations/Pair | 1.62 |
| Mean z-score | -0.87 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.67 |
| SVM RNA-class probability | 0.781585 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 9576226 94 - 22422827 GAAUAGUGGCUGGUCGGAUCAGCUGACCGGCGGAGCAGG--CGGUGGCAAUGGAGGUGGCGGCGCCUCCGGUGUAACCACAAAUCCCACAUCCGGC--- .....((((((((((((.....)))))))))(((....(--.(((.(((.((((((((....)))))))).))).))).)...)))))).......--- ( -43.80, z-score = -2.15, R) >droSim1.chrX 7626208 94 - 17042790 GAACAGUGGUUGGUCGGAUCAGCUGACCGGCGGAGCAGG--CGGUGGCAAUGGAGGUGGCGGCGCCUCCGGUGUAACCACAAACCCCACAUCCGGC--- .....((((((((((((.....)))))))))((.(...(--.(((.(((.((((((((....)))))))).))).))).)...)))))).......--- ( -38.70, z-score = -0.36, R) >droSec1.super_15 339973 96 - 1954846 GAACAGUGGUUGGUCGGAUCAGCUGAACGGCGGAGCAGGGCGGGUGGCAAUGGAGGUGGCGGCGCCUCCGGUGUAACCACAAACCCCACAUCCGGC--- ...((((((((.....))))).)))...(.((((...(((.(.((((((.((((((((....)))))))).))...))))...))))...)))).)--- ( -36.70, z-score = 0.20, R) >droYak2.chrX 18167482 94 - 21770863 GAACAGCGGCUGGUCGGAUCAGCUGACCGGCGGAGCAGG--CGGUGGCAAUGGAGGUGGCGGCGCCUCCGGUGUGACCACAAAUUCCACAUCCGGC--- ......(((((((((((.....)))))))))((((...(--.(((.(((.((((((((....)))))))).))).))).)...)))).....))..--- ( -44.70, z-score = -2.02, R) >droEre2.scaffold_4690 13206794 94 + 18748788 GAGCAGCGGCUGGUCGGAUCAGCUGACCGGCGGAGCAGG--CGGUGGCAAUGGAGGUGGCGGCGCCUCCGGUGUAACCACAAACUCCACAUCCGGC--- ......(((((((((((.....)))))))))((((...(--.(((.(((.((((((((....)))))))).))).))).)...)))).....))..--- ( -44.80, z-score = -1.81, R) >dp4.chrXL_group3a 801833 97 - 2690836 AAACAGCGGCUGGUCGGAUCAGCUGAGUGGCGGCGGCGG--CGGUGGCGCCUCGAAUGUUGGCGUCACAAACACAAACUCAACAGGCGGCGGCGGCGGA ...((.((((((((...))))))))..))((.((.((.(--(.((((((((.........)))))))).........((....)))).)).)).))... ( -39.80, z-score = -0.76, R) >droPer1.super_15 85866 97 + 2181545 AAACAGCGGCUGGUCGGAUCAGCUGAGUGGCGGCGGCGG--CGGUGGCGCCUCGAAUGUUGGCGUCACAAACACAAACUCAACAGGCGGCGGCGGCGGA ...((.((((((((...))))))))..))((.((.((.(--(.((((((((.........)))))))).........((....)))).)).)).))... ( -39.80, z-score = -0.76, R) >droMoj3.scaffold_6473 16099568 82 - 16943266 CAGCAGUGGCUGGUCGGAUCAGCUGAACACUGGUGGCGG--CGGCGGUGGCGGUGUUGGCGGCGUUGUAUCCACAGCCAGCGGC--------------- .....((.((((((.((((((((....(((((.((....--)).)))))((.((....)).)))))).))))...)))))).))--------------- ( -32.70, z-score = 0.34, R) >droGri2.scaffold_15203 65829 76 + 11997470 GAACAGCGGCUGGUCGGAUCAGCUAAACACU---GGCGG--CGGCGGUGGCGGUGUUAGUGUUGUUGCGACAGCAACCGGC------------------ .....(((((((((...))))))).((((((---((((.--((.(...).)).))))))))))(((((....)))))..))------------------ ( -28.40, z-score = -0.49, R) >consensus GAACAGCGGCUGGUCGGAUCAGCUGACCGGCGGAGCAGG__CGGUGGCAAUGGAGGUGGCGGCGCCUCCGGUGUAACCACAAACCCCACAUCCGGC___ ......((((((((...)))))))).................(((.(((..(((((((....)))))))..))).)))..................... (-19.09 = -18.49 + -0.60)


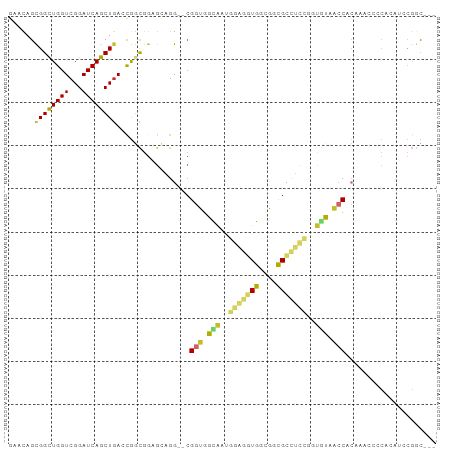
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:29:08 2011