
| Sequence ID | dm3.chrX |
|---|---|
| Location | 9,555,652 – 9,555,767 |
| Length | 115 |
| Max. P | 0.752170 |

| Location | 9,555,652 – 9,555,767 |
|---|---|
| Length | 115 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 65.68 |
| Shannon entropy | 0.63831 |
| G+C content | 0.43758 |
| Mean single sequence MFE | -29.02 |
| Consensus MFE | -8.93 |
| Energy contribution | -8.93 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.53 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.31 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.752170 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 9555652 115 - 22422827 GACCAUAUCAUUAUCGAGCCC--CCUUAAUCUG---GAUGUUUAUCUGUGCUGCAAAUCCCCAGCAUAAAGCCUGCAGGUGGGGGGGACUCGAAAAUCUUUAGCACUUCAAAAGUUCUUA .............((((((((--(((((.((((---.(.((((...(((((((........))))))))))).).)))))))))))).)))))........................... ( -36.50, z-score = -2.47, R) >droGri2.scaffold_15203 44050 99 + 11997470 ------UUCA-CACUGCAGGUGUUCCUGAUAGGAAAUAAACCUUUAUGGUCGACGAUAAUCAAGAUUAAAGUCGUUAGCUGAGGGUGAAUGGGAA----UUAAGUUUACA---------- ------((((-(.(((((((....)))).)))........((((..(((.((((..((((....))))..)))))))...)))))))))......----...........---------- ( -23.50, z-score = -1.21, R) >droEre2.scaffold_4690 13186061 100 + 18748788 AAUCAUAUCA-UAUCAGUC----CUUUAAUCUG---GGGGCUUAUCGGCGCUGCAAAUGCCCAGCAUAAAACCUGCAGGUGGGGA-AAGUCGAAAAUCUGUAACUC--CAA--------- ..........-....((((----(((......)---))))))..(((((((((........))))......((((....))))..-..))))).............--...--------- ( -21.50, z-score = 0.65, R) >droYak2.chrX 18146316 114 - 21770863 AUCCGUAUCA-UAUCGGGCUCCAUUUUAAUCUA--CUAGACUUAUCGUUGCUGCAAAUCCCCAGCAGAAAACCCGCAGGUGGGGA---GUCCAAAAUCUGUAGCUCUUCAAAAGUUGUUA ..........-....(((((((........(((--((.(........((((((........))))))........).))))))))---))))........(((((.......)))))... ( -23.39, z-score = -0.05, R) >droSec1.super_15 319259 111 - 1954846 GACCAUAUCA-UAUCGAGCCC--CCUUAAUCUG---UAUGCUUAUCUGUGCUGCAAAUCCCCAGCAUAAAGCCUGCAGGUGGGGA---CUCGAAAAUCUCUAGCUCUUCAAAAGUUCUUA ..........-..(((((.((--((...(((((---((.((((...(((((((........))))))))))).))))))))))).---)))))........................... ( -34.60, z-score = -4.21, R) >droSim1.chrX_random 2755778 111 - 5698898 GACCAUAUCA-UAUCGAGCCC--CCUUAAUCUG---UAUGCUUAUCUGUGCUGCAAAUCCCCAGCAUAAAGCCUGCAGGUGGGGA---CUCGAAAAUCUCUAGCUCUUCAAAAGUUCUUA ..........-..(((((.((--((...(((((---((.((((...(((((((........))))))))))).))))))))))).---)))))........................... ( -34.60, z-score = -4.21, R) >consensus GACCAUAUCA_UAUCGAGCCC__CCUUAAUCUG___UAUGCUUAUCUGUGCUGCAAAUCCCCAGCAUAAAGCCUGCAGGUGGGGA___CUCGAAAAUCUCUAGCUCUUCAAAAGUUCUUA .............(((((.....((((...................(((((((........)))))))..(((....)))))))....)))))........................... ( -8.93 = -8.93 + 0.01)
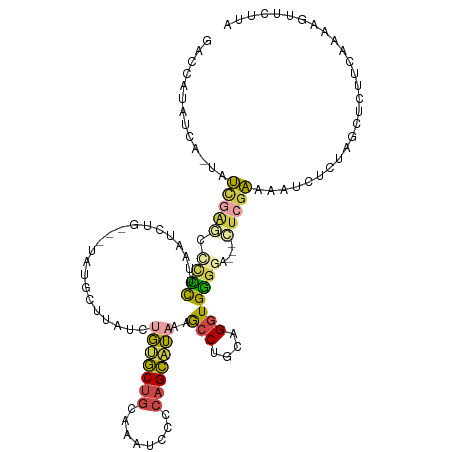


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:29:06 2011