
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 10,761,501 – 10,761,591 |
| Length | 90 |
| Max. P | 0.830195 |

| Location | 10,761,501 – 10,761,591 |
|---|---|
| Length | 90 |
| Sequences | 9 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 56.88 |
| Shannon entropy | 0.87133 |
| G+C content | 0.55290 |
| Mean single sequence MFE | -27.51 |
| Consensus MFE | -12.26 |
| Energy contribution | -12.48 |
| Covariance contribution | 0.22 |
| Combinations/Pair | 1.89 |
| Mean z-score | -0.51 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.830195 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 10761501 90 + 23011544 ---CGAAAUGAUUGCCAACUUGGCGUUGCCUUUCCUGCUGAUGAUCGCGGAAGCAGCAGC-CACCGCGG------AGCAGGUGAAGUACCUGGAGUACCU ---.((((.(..((((.....))))...).))))(((((.....((((((..((....))-..))))))------))))).....((((.....)))).. ( -26.70, z-score = 0.08, R) >droSim1.chr2L 10561078 93 + 22036055 CGAAAUAAUGAUUGCCAACUUGGCAUUGCCUUUCCUGCUGAUCAUCGCGGAAGCAGCAGC-CACCGCGG------AGCAGGUGGAGUACCUGGAAUACCU ............((((.....))))...((...((((((.....((((((..((....))-..))))))------)))))).))................ ( -27.60, z-score = -0.28, R) >droSec1.super_3 6180034 93 + 7220098 CGAAAUAAUGAUUGCCAACUUGGCAUUGCCUUUCCUGCUGAUCAUCGCGGAAGCAGCAGC-CACCGCGG------AGCAGGUGGAGUACCUGGAGUACCU ............((((.....))))...((...((((((.....((((((..((....))-..))))))------)))))).)).((((.....)))).. ( -29.70, z-score = -0.59, R) >droYak2.chr2L 7166757 90 + 22324452 ---CAAAAUGAUUGCCAUGUUGGCUUUGCCCUUCCUGCUGAUGAUCGCGGAAGCAGCAGC-CACCGCGG------AGCAGGUGGAGUACCUGGAGUACCU ---..........(((.....))).....((..((((((.....((((((..((....))-..))))))------)))))).)).((((.....)))).. ( -28.70, z-score = -0.13, R) >droEre2.scaffold_4929 11971607 81 - 26641161 ---CGCAAUGAUUGCCAACCUGGCGUUGCCCUUCCUGCUGAUGAUCGCGGAAGCAGCAGC-CACCGCGG------AGCAGGUGGAGUACCU--------- ---.(((((....(((.....))))))))((..((((((.....((((((..((....))-..))))))------)))))).)).......--------- ( -30.20, z-score = -0.91, R) >droAna3.scaffold_12916 14612584 87 + 16180835 ---UGAAAUGAUAUCCAAUUUGGCGCUAAUUCCGCUGCUAAUGAUUGUGGGAGCAGCAGC-CACCGCAG------AGCAGGUGGAGUACCUCAAUCC--- ---(((..((...(((.....((((((..((((((..(....)...))))))..))).))-)(((....------....)))))).))..)))....--- ( -23.80, z-score = -0.11, R) >droPer1.super_10 1980104 93 + 3432795 ---UCAAAUGUUUUCCAAUCUGCCUCUGGCUGCGCUG-CUGCUCCCGCUGCCACUGCUGCUCGUCGCGGUG--CAGGCAAC-GGAGUAUGUGAAUCCGCC ---......(..(((((..((.((..((.((((((((-(.((....((.((....)).))..)).))))))--))).))..-))))..)).)))..)... ( -29.40, z-score = -0.27, R) >droWil1.scaffold_180703 294322 76 - 3946847 ---------UUUGGCCAUUACAACUAUAAUUAUUCUCAUAAUAGAUGCAGUGGCAGCUGCUUCUGGUCUUG----GACAACCUGAAAUU----------- ---------.(((.(((..((........((((....))))((((.(((((....))))).))))))..))----).))).........----------- ( -14.60, z-score = -0.18, R) >droMoj3.scaffold_6500 13471170 80 - 32352404 ---UUAAGCUGUUGCCG------CUCCAGCUGCUCCAUCUGCUCCAGCUGCCGCUGCUCCUCAUGGCAGCUGCCGGGCAGCCGGAGUAC----------- ---...(((((......------...)))))(((((..(((((((((((((((.((.....)))))))))))..))))))..)))))..----------- ( -36.90, z-score = -2.14, R) >consensus ___CAAAAUGAUUGCCAACUUGGCUUUGCCUUUCCUGCUGAUGAUCGCGGAAGCAGCAGC_CACCGCGG______AGCAGGUGGAGUACCUGGA_UA_C_ .............................((((((((((.....((((((..((....))...))))))......)))))).)))).............. (-12.26 = -12.48 + 0.22)
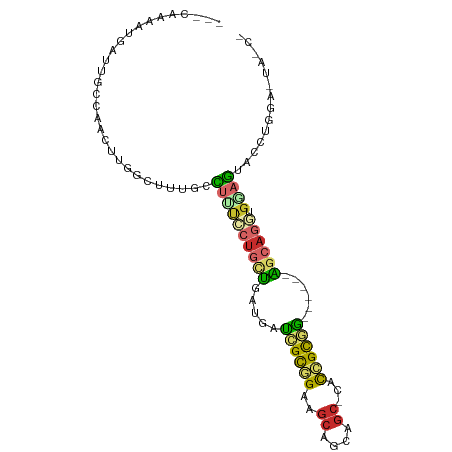


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:30:54 2011