| Sequence ID | dm3.chrX |
|---|---|
| Location | 9,399,321 – 9,399,417 |
| Length | 96 |
| Max. P | 0.500000 |

| Location | 9,399,321 – 9,399,417 |
|---|---|
| Length | 96 |
| Sequences | 7 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 60.09 |
| Shannon entropy | 0.73837 |
| G+C content | 0.40827 |
| Mean single sequence MFE | -24.53 |
| Consensus MFE | -6.84 |
| Energy contribution | -8.89 |
| Covariance contribution | 2.05 |
| Combinations/Pair | 1.60 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.28 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
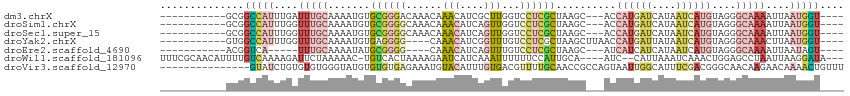
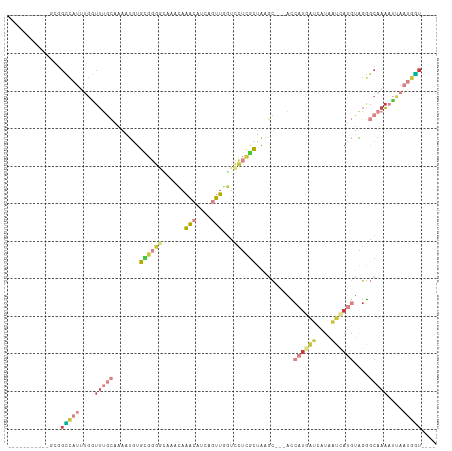
>dm3.chrX 9399321 96 - 22422827 -----------GCGGCCAUUUGAUUUGCAAAAUGUGCGGGACAAACAAACAUCGCUUGGUCCUCGCUAAGC---ACCAUGAUCAUAAUCAUGUAGGGCAAAAUUAAUGGU---- -----------...((((((...(((((...((((...(......)..)))).(((((((....)))))))---..((((((....))))))....)))))...))))))---- ( -25.40, z-score = -0.96, R) >droSim1.chrX 7494085 96 - 17042790 -----------GCGGCCAUUUGGUUUGCAAAAUGUGCGGGGCAAACAAACAUCAGUUGGUCCUCGCUAAGC---ACCAUGAUCAUAAUCAUGUAGGGCAAAAUUAAUGGU---- -----------...((((((...(((((.......(((((((.(((........))).)))).))).....---..((((((....))))))....)))))...))))))---- ( -28.60, z-score = -1.64, R) >droSec1.super_15 165738 96 - 1954846 -----------GCGGCCAUUUGGUUUGCAAAAUGUGCGGGGCAAACAAACAUCAGUUGGUCCUCGCUAAGC---ACCAUGAUCAUAAUCAUGUAGGGCAAAAUUAAUGGU---- -----------...((((((...(((((.......(((((((.(((........))).)))).))).....---..((((((....))))))....)))))...))))))---- ( -28.60, z-score = -1.64, R) >droYak2.chrX 17991396 95 - 21770863 -----------GUGGCCAUUUGGUUUGCAAAAUGUGAGGGG----CAAACAUCGGUUUGUCCUCGCUAAGCUUAACCAUGAUUAUAAUCAUGUAGGGCAAACUUAAUGGU---- -----------...((((((.(((((((.....(((((((.----(((((....))))))))))))....((((..((((((....))))))))))))))))).))))))---- ( -37.50, z-score = -4.94, R) >droEre2.scaffold_4690 13033609 87 + 18748788 -----------ACGGUCA-----UUUGCAAAAUAUGCGGGG----CAAACAUCAGUUUGUCCUCGCUAAGC---AUCAUCAUCAUAAUCAUGUAGGGCAAAAUUAAUAGU---- -----------.......-----(((((.......((((((----(((((....)))))))).)))...((---((.((.......)).))))...))))).........---- ( -20.70, z-score = -2.99, R) >droWil1.scaffold_181096 9766506 104 + 12416693 UUUCGCAACAUUUUGUCAAAAGAUUCUAAAAAC-UGUCACUAAAAGAAUCAUCAAAUUUUUUCCAUUGCA----AUC--CAUUAAAUCAAACUGGAGCCUAAUUAAGGAUA--- ....((((...(((....)))((((((......-..........))))))...............)))).----.((--((...........)))).(((.....)))...--- ( -10.19, z-score = 0.43, R) >droVir3.scaffold_12970 6036906 99 - 11907090 ---------------GUAUCUGUGUGUGGGUAUGUGUGUGAGAAAUGUACAUUUGUGACGUUUUGCAACCGCCAGUAAUUGGCAUUUCGACGGGCAACAAGAACAAAACUGUUU ---------------((.(((.(((.((((((((((..(.....)..))))).((..(....)..)))))(((((...))))).......)).)))...))))).......... ( -20.70, z-score = 1.42, R) >consensus ___________GCGGCCAUUUGGUUUGCAAAAUGUGCGGGGCAAACAAACAUCAGUUGGUCCUCGCUAAGC___ACCAUGAUCAUAAUCAUGUAGGGCAAAAUUAAUGGU____ ..............(((((....(((((.......((((((......(((....)))...))))))..........((((((....))))))....)))))....))))).... ( -6.84 = -8.89 + 2.05)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:28:29 2011