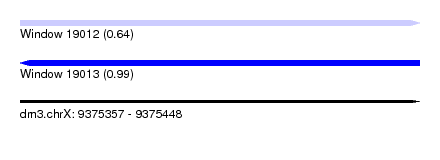

| Sequence ID | dm3.chrX |
|---|---|
| Location | 9,375,357 – 9,375,448 |
| Length | 91 |
| Max. P | 0.990916 |

| Location | 9,375,357 – 9,375,448 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 74.77 |
| Shannon entropy | 0.40824 |
| G+C content | 0.36726 |
| Mean single sequence MFE | -18.15 |
| Consensus MFE | -13.06 |
| Energy contribution | -13.25 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.08 |
| Structure conservation index | 0.72 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.641716 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
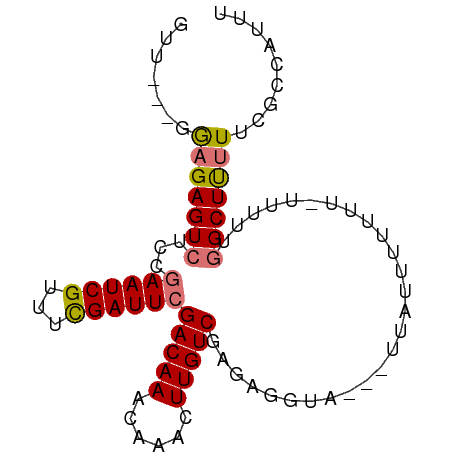
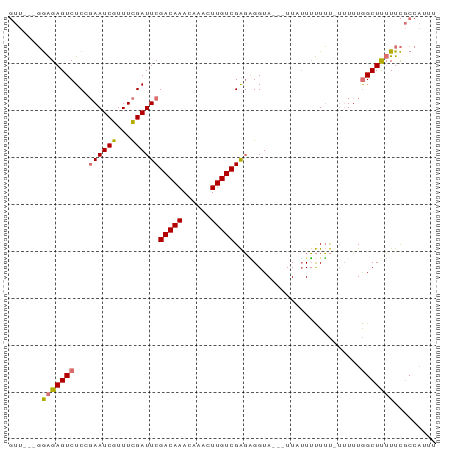
>dm3.chrX 9375357 91 + 22422827 GUUUGAGGAGAGUCUCGGAAUCGUUUCGAUUCGACAAACAAACUUGUCGAGGGGUAAAAUUUUUUUUUUGUUUUUGGCUUUUUCGCCAAUU ....((((((((((((.((((((...))))))(((((......)))))))).......)))))))))......(((((......))))).. ( -20.11, z-score = -0.22, R) >droEre2.scaffold_4690 13010004 74 - 18748788 -UUU--UAGGAGUCUC--AAUCGUUUUGAUUCGACAAAC----UUGUCGGUUGGUAUUAUUUGUACUGC-UUUUUGGCUCGUUU------- -...--...(((((((--((.....)))).((((((...----.))))))..(((((.....)))))..-.....)))))....------- ( -15.70, z-score = -1.77, R) >droSec1.super_15 142555 85 + 1954846 GUU---GGAGAGUCUCCGAAUCGUUUCGAUUCGACAAACAAACUUGUCGAGAGGUA---UUAUUUUUUUAUUUUUGGCUUUUUCGCCAUUU ...---((((...))))((((((...))))))(((((......)))))((((((..---....)))))).....((((......))))... ( -19.60, z-score = -1.44, R) >droSim1.chrX 7472425 82 + 17042790 GUU---GGAGAGUCUCCGAAUCGUUUCGAUUCGACAAACAAACUUGUCGAGAGGUA---UUAUUUUUUU---UUUAGCUUUUUCGCCAUUU (((---((((((((((.((((((...))))))(((((......)))))))))....---.......)))---))))))............. ( -17.20, z-score = -0.90, R) >consensus GUU___GGAGAGUCUCCGAAUCGUUUCGAUUCGACAAACAAACUUGUCGAGAGGUA___UUAUUUUUUU_UUUUUGGCUUUUUCGCCAUUU .......(((((((...((((((...))))))(((((......)))))...........................)))))))......... (-13.06 = -13.25 + 0.19)

| Location | 9,375,357 – 9,375,448 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 91 |
| Reading direction | reverse |
| Mean pairwise identity | 74.77 |
| Shannon entropy | 0.40824 |
| G+C content | 0.36726 |
| Mean single sequence MFE | -16.98 |
| Consensus MFE | -10.45 |
| Energy contribution | -12.32 |
| Covariance contribution | 1.88 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.59 |
| Structure conservation index | 0.62 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.45 |
| SVM RNA-class probability | 0.990916 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
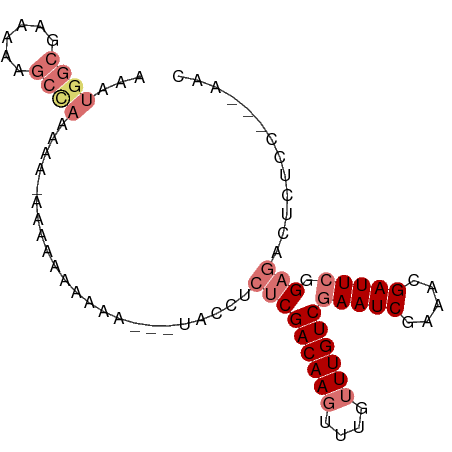
>dm3.chrX 9375357 91 - 22422827 AAUUGGCGAAAAAGCCAAAAACAAAAAAAAAAUUUUACCCCUCGACAAGUUUGUUUGUCGAAUCGAAACGAUUCCGAGACUCUCCUCAAAC ..(((((......)))))......................(((((((((....))))))((((((...)))))).)))............. ( -20.00, z-score = -3.05, R) >droEre2.scaffold_4690 13010004 74 + 18748788 -------AAACGAGCCAAAAA-GCAGUACAAAUAAUACCAACCGACAA----GUUUGUCGAAUCAAAACGAUU--GAGACUCCUA--AAA- -------....(((.(.....-....................(((((.----...)))))((((.....))))--..).)))...--...- ( -8.60, z-score = -1.29, R) >droSec1.super_15 142555 85 - 1954846 AAAUGGCGAAAAAGCCAAAAAUAAAAAAAUAA---UACCUCUCGACAAGUUUGUUUGUCGAAUCGAAACGAUUCGGAGACUCUCC---AAC ...((((......))))...............---....((((((((((....))))))((((((...)))))).))))......---... ( -20.80, z-score = -3.45, R) >droSim1.chrX 7472425 82 - 17042790 AAAUGGCGAAAAAGCUAAA---AAAAAAAUAA---UACCUCUCGACAAGUUUGUUUGUCGAAUCGAAACGAUUCGGAGACUCUCC---AAC ...((((......))))..---..........---....((((((((((....))))))((((((...)))))).))))......---... ( -18.50, z-score = -2.56, R) >consensus AAAUGGCGAAAAAGCCAAAAA_AAAAAAAAAA___UACCUCUCGACAAGUUUGUUUGUCGAAUCGAAACGAUUCGGAGACUCUCC___AAC ...((((......)))).......................(((((((((....))))))(((((.....))))).)))............. (-10.45 = -12.32 + 1.88)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:28:24 2011