
| Sequence ID | dm3.chrX |
|---|---|
| Location | 9,309,527 – 9,309,662 |
| Length | 135 |
| Max. P | 0.824335 |

| Location | 9,309,527 – 9,309,662 |
|---|---|
| Length | 135 |
| Sequences | 5 |
| Columns | 164 |
| Reading direction | reverse |
| Mean pairwise identity | 71.59 |
| Shannon entropy | 0.49478 |
| G+C content | 0.54419 |
| Mean single sequence MFE | -37.25 |
| Consensus MFE | -21.38 |
| Energy contribution | -23.18 |
| Covariance contribution | 1.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.57 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.81 |
| SVM RNA-class probability | 0.824335 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 9309527 135 - 22422827 -----------------UCCCGCCUGCCGUUCCUCAAUUUCCCAUUUUCCCCCGCCACGUUCCCAGCAUUUUGUAGCCCAACUCGGGCG------------AACUGCAAUCGCCGGGGCUUAUUCACUUGUGAGUAAAAGUUGGCCAGAGCCAAAAGCAAACUU -----------------.(((((.(((.((((.....................((..........))........((((.....)))))------------))).)))...).))))(((((((((....)))))))...(((((....)))))..))...... ( -33.90, z-score = -0.42, R) >droSim1.chrX 7409356 135 - 17042790 -----------------UCCUCCCGCCCGGCG-----UUCCCCAAUUCCCC-------GUUCCCAGCAUUUUGUAGCCCAACCCGGGCGAACCCCGCAUUCAACUGCAAUCGCCGGGGCUUAUUCACUUGUGAGUAAAAGUUGGCCAGAGCCAAAAGCAAACUU -----------------........(((((((-----..............-------((((...((.....)).((((.....))))))))...(((......)))...)))))))(((((((((....)))))))...(((((....)))))..))...... ( -40.70, z-score = -2.19, R) >droSec1.super_15 79254 135 - 1954846 -----------------UCCUCCCGCCCGGCG-----UUCCCCAUUUUUUC-------CUUCCCAGCAGUUUGUAGCCCAACUCGGGCGAACCCCGCAUUCAACUGCAAUCGCCGGGGCUUAUUCACUUGUGAGUAAAAGUUGGCCAGAGCCAAAAGCAAACUU -----------------........(((((((-----.....)........-------.......((((((....((((.....))))(((.......)))))))))....))))))(((((((((....)))))))...(((((....)))))..))...... ( -39.70, z-score = -1.67, R) >droYak2.chrX 17898261 160 - 21770863 -UCCCGCCUGCCGUUCCCCAAUUUCCCAUAUUCCCAUUUUCCCCUUUUCCACCAGCACAUUCCCUGCAUUUUGCAGCCCAAGCCCCGCAUUCAACGCAUUCAA---CACUCGCCGGGGCUUUUUCACUUGUGAGUAAAAGUUGGCCAGAGCCAAAAGCAAACUA -.......(((................................(((((((((.((........((((.....))))...(((((((((...............---.....).)))))))).....)).))).).)))))(((((....)))))..)))..... ( -30.25, z-score = -0.90, R) >droEre2.scaffold_4690 12942388 161 + 18748788 ACCCGCACACCCGCUCACCCGCUCACCCGGUGCCCAAUUUCCCAUUUUCCC--AGCCAGUUCCCAGCAUUUUCCAGCCCAAGCCAGGCGAACCCCGCAUUCAACGGCAAUCGCC-GGGCUUAUUCACUUGUGAGCAAAAGUUGGCCAGAGCCAAAAGCAAACUU ............(((.....((((((..((((...................--............((........))..(((((.(((((...(((.......)))...)))))-.)))))...)))).)))))).....(((((....))))).)))...... ( -41.70, z-score = -1.85, R) >consensus _________________UCCUCCCGCCCGGUGC_CA_UUUCCCAUUUUCCC____C__GUUCCCAGCAUUUUGUAGCCCAACCCGGGCGAACCCCGCAUUCAACUGCAAUCGCCGGGGCUUAUUCACUUGUGAGUAAAAGUUGGCCAGAGCCAAAAGCAAACUU .............................................................(((.((........((((.....)))).......((........))....)).)))(((((((((....)))))))...(((((....)))))..))...... (-21.38 = -23.18 + 1.80)

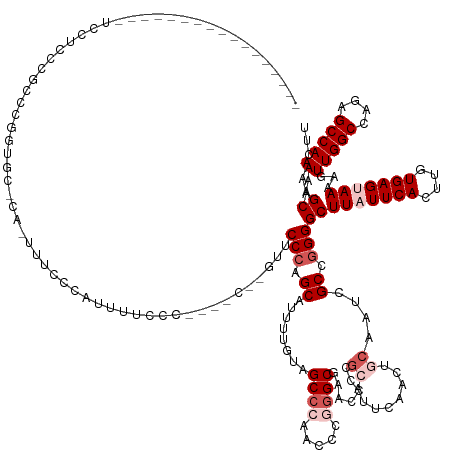
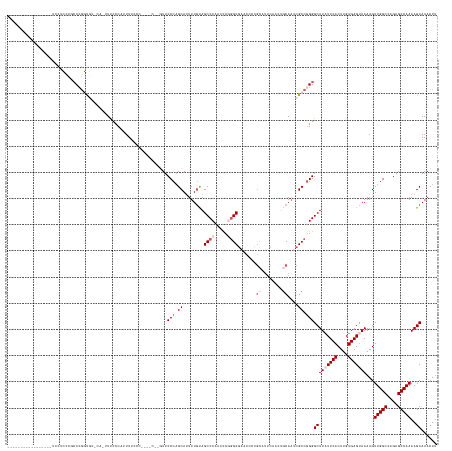
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:28:02 2011