
| Sequence ID | dm3.chrX |
|---|---|
| Location | 9,021,613 – 9,021,714 |
| Length | 101 |
| Max. P | 0.928700 |

| Location | 9,021,613 – 9,021,714 |
|---|---|
| Length | 101 |
| Sequences | 7 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 67.97 |
| Shannon entropy | 0.60159 |
| G+C content | 0.47863 |
| Mean single sequence MFE | -17.93 |
| Consensus MFE | -9.85 |
| Energy contribution | -10.03 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.22 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.34 |
| SVM RNA-class probability | 0.928700 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 9021613 101 - 22422827 -GCAUGCUCAUCCUCAUCGUCCUGUCACGGCAAUCCUCCUCAUUAUCCUCAUUGCCGGCAGCGUGAAAUUUUCAAUUCGCCAUAGACGAACACAAACGUUUG -...............(((((((((..(((((((................))))))))))).(((((........)))))....)))))............. ( -19.79, z-score = -1.44, R) >dp4.chrXL_group1e 5562132 77 + 12523060 -------------------AUUUGCCAUGGCAAUCCCCCUCAUUAUCCUCAUUGCCAGGAGCGUGAAAUUUUCAAUUCUCCACAGGCCAACGUUUG------ -------------------....(((.(((((((................)))))))((((..(((.....)))...))))...))).........------ ( -18.39, z-score = -2.30, R) >droPer1.super_13 1593415 77 - 2293547 -------------------AUUUGCCAUGGCAAUCCCCCUCAUUAUCCUCAUUGCCAGGAGCGUGAAAUUUUCAAUUCUCCACAGGCCAACGUUUG------ -------------------....(((.(((((((................)))))))((((..(((.....)))...))))...))).........------ ( -18.39, z-score = -2.30, R) >droSec1.super_34 381905 89 - 457129 -------UCAGCUU------CCUGUCACGGCAAUCCUCCUCAUUAUCCUCAUUGCCGGCAGCGUGAAAUUUUCAAUUCGCCAUAGACGAACACAAACGUUUG -------.......------...(((..((((((................))))))(((....(((.....)))....)))...)))((((......)))). ( -16.49, z-score = -1.13, R) >droYak2.chrX 9476106 98 + 21770863 -GCAUGCUCAUCCU---CGUCCUGUCACGGCAAUCCUCCUCAUUAUCCUCAUUGCCGGGAGCGUGAAAUUUUCAAUUCGCCAUAGACGAACACAAACGUUUG -.(((((((.....---.(......).(((((((................))))))).))))))).................((((((........)))))) ( -20.19, z-score = -1.01, R) >droEre2.scaffold_4690 12652352 98 + 18748788 -UCAUGCUCAGCCU---CGUCCUGUUACGGCAAUCCUCCUCAUUAUCCUCAUUGCCGGGAGCGUGAAAUUUUCAAUUCGCCAUAGACGAACACAAACGUUUG -(((((((((((..---......))).(((((((................))))))).))))))))................((((((........)))))) ( -22.29, z-score = -1.66, R) >droVir3.scaffold_12932 1885378 90 - 2102469 ACAGUCGCCGUCGUCGUCGCCAUGUCCAGUUAGUUCACAGUUUCAUCUCC--UGCCGACAGCGUGAAAAUUUCAAUUCGCCAUACAACUUUG---------- ......((.((((.........((..(.....)..))(((.........)--)).)))).))(((((........)))))............---------- ( -10.00, z-score = 1.32, R) >consensus ____U__UCAUCCU___CGUCCUGUCACGGCAAUCCUCCUCAUUAUCCUCAUUGCCGGGAGCGUGAAAUUUUCAAUUCGCCAUAGACGAACACAAACGUUUG .......................(((.(((((((................)))))))...(.(((((........))))))...)))............... ( -9.85 = -10.03 + 0.19)
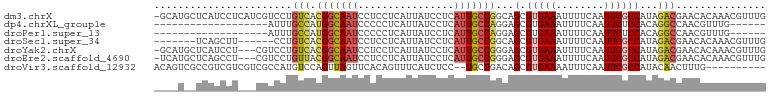
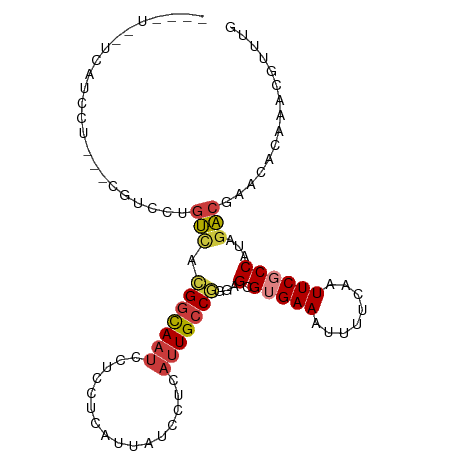
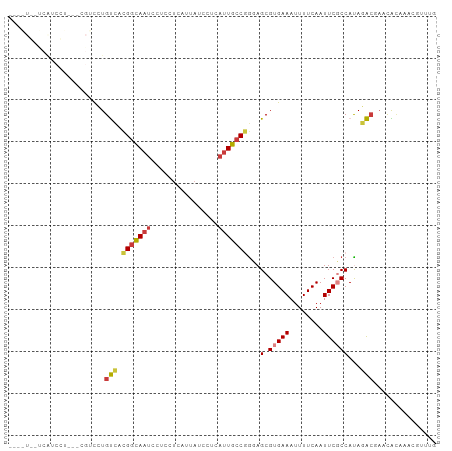
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:27:22 2011