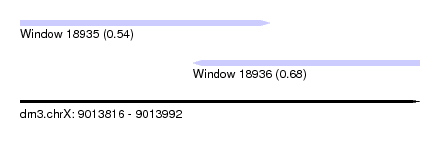
| Sequence ID | dm3.chrX |
|---|---|
| Location | 9,013,816 – 9,013,992 |
| Length | 176 |
| Max. P | 0.677993 |

| Location | 9,013,816 – 9,013,926 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 74.64 |
| Shannon entropy | 0.41107 |
| G+C content | 0.47699 |
| Mean single sequence MFE | -23.98 |
| Consensus MFE | -16.30 |
| Energy contribution | -16.43 |
| Covariance contribution | 0.13 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.08 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.538489 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
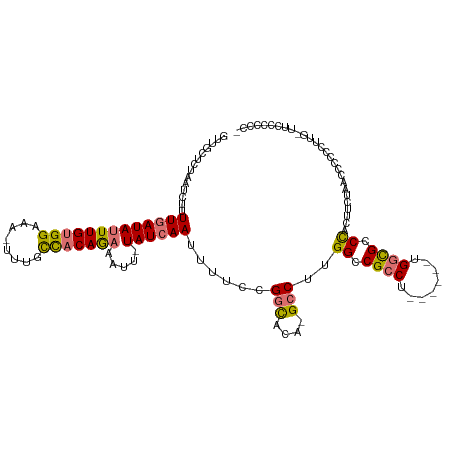
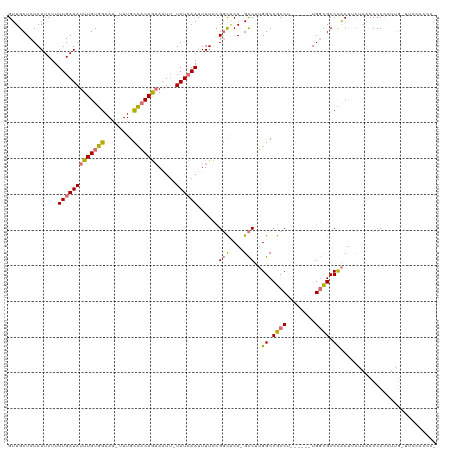
>dm3.chrX 9013816 110 + 22422827 GUUGCUCUAAUCUUUUGAUAUUUGUGGAAA-UUUGCCACAGAAAUU-UAUCAAUUUUCCGGCACA-GCCUUGGCCGCCU------UGGCGCCCACUUCGAACCCCCCUUGAUUCCCCAC- ((((..((......(((((((((((((...-....)))))))....-))))))......))..))-))..(((.(((..------..))).))).........................- ( -22.20, z-score = -0.86, R) >droSec1.super_34 374092 110 + 457129 GUUGCUCUAAUCUUUUGAUAUUUGUGGAAA-UUUGCCACAGAAAUU-UAUCAAUUUUCCGGCACA-GCCUUGGCCGCCU------UGGCGCCCACUUAAAACCCCCCUUGUUUCCACCC- .........(((....)))....(((((((-(.((((...((((((-....))))))..))))..-....(((.(((..------..))).)))...............))))))))..- ( -23.50, z-score = -1.48, R) >droYak2.chrX 9468388 96 - 21770863 GUUGCUCUAAUCUUUUGAUAUUUGUGGAAA-UUUGCCACAGAAAUU-UAUCAAUUUUCCGGCUCA-GCCUUGGCCGCCU------UGGCGCCCACUUCUCCGCCC--------------- ..............(((((((((((((...-....)))))))....-))))))......(((..(-(...(((.(((..------..))).)))...))..))).--------------- ( -23.70, z-score = -1.61, R) >dp4.chrXL_group1e 5554501 120 - 12523060 GUUGCUCUAAUCUUUUGAUAUUUGUGGAAAAUUUGUUUCAAUAUUGAUAUGAAUUUUGUGCUGCAAGUCGAGGCCGCCUGCCGCCUGCUGCCUGCUGCUAACCCCCGUGGCACUACGCCC ((..(....(((....)))......((((((((..(.(((....))).)..))))))..((.(((.((.(((((.(....).)))).).)).))).))......)))..))......... ( -26.50, z-score = -0.39, R) >consensus GUUGCUCUAAUCUUUUGAUAUUUGUGGAAA_UUUGCCACAGAAAUU_UAUCAAUUUUCCGGCACA_GCCUUGGCCGCCU______UGGCGCCCACUUCUAACCCCCCUUG_UUCCCCCC_ ..............(((((((((((((........))))))).....))))))......(((....)))..((.((((........)))).))........................... (-16.30 = -16.43 + 0.13)

| Location | 9,013,892 – 9,013,992 |
|---|---|
| Length | 100 |
| Sequences | 3 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 73.44 |
| Shannon entropy | 0.35668 |
| G+C content | 0.60758 |
| Mean single sequence MFE | -34.53 |
| Consensus MFE | -22.24 |
| Energy contribution | -22.30 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.64 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.677993 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
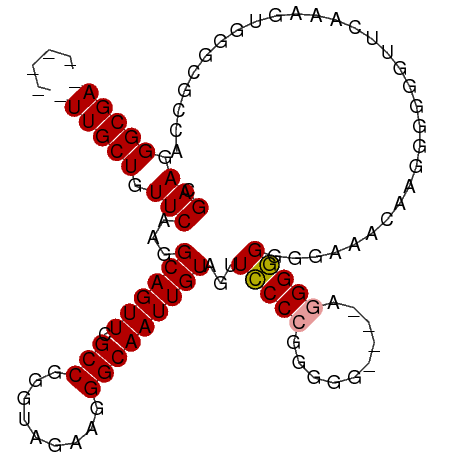
>dm3.chrX 9013892 100 - 22422827 CGAAGGGCGA------UUGCUGUUCAAGGCAGUUCGCCGAGUAGAAGGGCAAUUGUAGUUACCCGGGGG----AGGGUGGGGAAUCAAGGGGGGUUCGAAGUGGGCGCCA .((...((((------(((((.(((..(((.....))).....))).)))))))))..((((((.....----.))))))....))....((.((((.....)))).)). ( -33.80, z-score = -1.93, R) >droSec1.super_34 374168 110 - 457129 CGAAGGGCGAUUGCUGUUGCUGUUCAAGGCAGUUCGCCGGGUAGACGGGCAAUUGUAGUUCCCCGGGGGGUGUGGGGGGUGGAAACAAGGGGGGUUUUAAGUGGGCGCCA .....((((.(..((...(((.(((...((((((.(((.(.....).))))))))).(..(((((.......)))))..)(....)..))).)))....))..).)))). ( -39.40, z-score = -1.21, R) >droYak2.chrX 9468464 88 + 21770863 CGAAGGGCGA------UUGCUGUUCAAGGCAGUUUGCCGGGUAGAAGGGCAAUUGUAGUUCCCAAGGGG----AGGGGGGGCGGAGAAGUGGGCGCCA------------ .....((((.------(..((.(((...((((((.(((.........))))))))).((((((......----...)))))).))).))..).)))).------------ ( -30.40, z-score = -1.78, R) >consensus CGAAGGGCGA______UUGCUGUUCAAGGCAGUUCGCCGGGUAGAAGGGCAAUUGUAGUUCCCCGGGGG____AGGGGGGGGAAACAAGGGGGGUUCAAAGUGGGCGCCA .(((.((((((....)))))).)))...((((((.(((.........)))))))))...(((((..........)))))............................... (-22.24 = -22.30 + 0.06)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:27:21 2011