
| Sequence ID | dm3.chrX |
|---|---|
| Location | 8,914,360 – 8,914,454 |
| Length | 94 |
| Max. P | 0.999279 |

| Location | 8,914,360 – 8,914,454 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 82.39 |
| Shannon entropy | 0.28632 |
| G+C content | 0.43132 |
| Mean single sequence MFE | -25.48 |
| Consensus MFE | -20.61 |
| Energy contribution | -21.30 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.08 |
| Mean z-score | -3.46 |
| Structure conservation index | 0.81 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.34 |
| SVM RNA-class probability | 0.998379 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
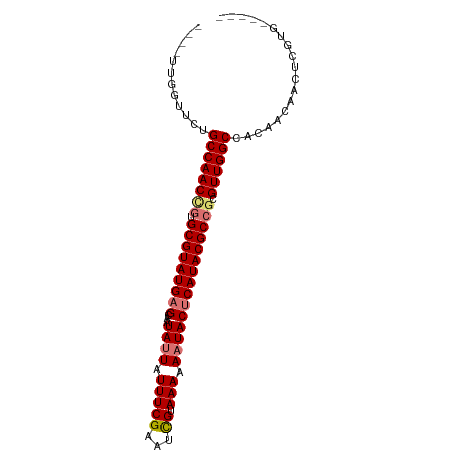
>dm3.chrX 8914360 94 + 22422827 GUUAUUGUUUCUGCCAACCGUGCGUAUGAGCAAUAUUAUUUCGAAUCGUAAAAAUUACUCAUACGCCGCGUUGGCCACAACAACUGAAAGCGUG (((.((((....((((((((.(((((((((.(((.((((........))))..))).))))))))))).)))))).)))).))).......... ( -29.20, z-score = -3.64, R) >droYak2.chrX 9372227 89 - 21770863 AAUUUCGGUUCUGCCAACCAUGCGUAUGAGCAAUAUUAUUUCGAAUUGCAAAAAAUACGCAUACGCCUCGUUGGCCACAACAACUACUG----- .......(((..((((((...(((((((.(((((..........)))))..........)))))))...))))))...)))........----- ( -21.20, z-score = -2.38, R) >droSim1.chrX_random 2621009 78 + 5698898 --------UUCUGCCAACCGUGCGUAUGAGCAAUAUUAUUUCGAAUCGUAAAAAAUACUCAUACGCCGCGUUGGCCACAAC---UCGUG----- --------....((((((((.(((((((((...((((.(((((...)).))).))))))))))))))).))))))(((...---..)))----- ( -26.60, z-score = -4.79, R) >droSec1.super_34 276927 85 + 457129 ----UUGGUUCUGCCAACUGUGCGUAUGAGCAAUAUUAUUUCGAAUCGUAAAAAAUACUCAUACGCCGCGUUGGCCACAACAACUCGUG----- ----...(((..((((((((.(((((((((...((((.(((((...)).))).))))))))))))))).))))))...)))........----- ( -24.90, z-score = -3.04, R) >consensus ____UUGGUUCUGCCAACCGUGCGUAUGAGCAAUAUUAUUUCGAAUCGUAAAAAAUACUCAUACGCCGCGUUGGCCACAACAACUCGUG_____ ............((((((((.(((((((((...((((.(((((...)).))).))))))))))))))).))))))................... (-20.61 = -21.30 + 0.69)


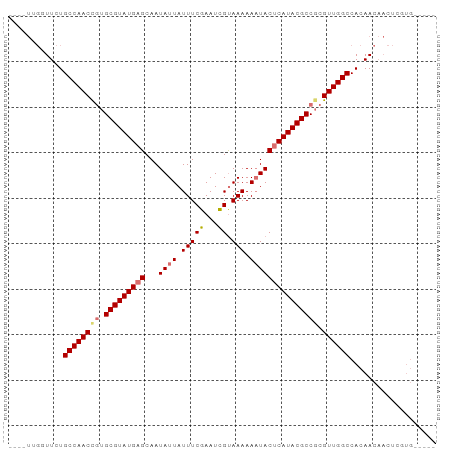
| Location | 8,914,360 – 8,914,454 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 82.39 |
| Shannon entropy | 0.28632 |
| G+C content | 0.43132 |
| Mean single sequence MFE | -31.35 |
| Consensus MFE | -25.60 |
| Energy contribution | -25.85 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -4.10 |
| Structure conservation index | 0.82 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.76 |
| SVM RNA-class probability | 0.999279 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
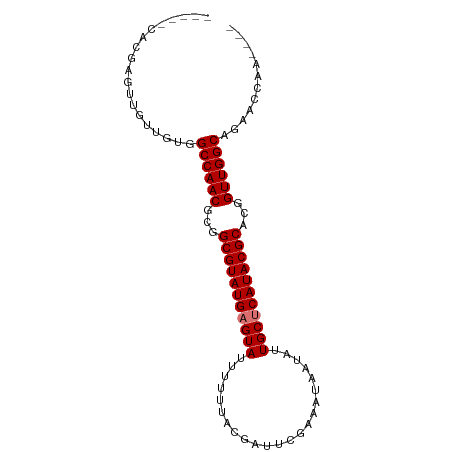
>dm3.chrX 8914360 94 - 22422827 CACGCUUUCAGUUGUUGUGGCCAACGCGGCGUAUGAGUAAUUUUUACGAUUCGAAAUAAUAUUGCUCAUACGCACGGUUGGCAGAAACAAUAAC ..........((((((((.((((((.(((((((((((((((.(((.((...)))))....))))))))))))).))))))))....)))))))) ( -38.70, z-score = -6.16, R) >droYak2.chrX 9372227 89 + 21770863 -----CAGUAGUUGUUGUGGCCAACGAGGCGUAUGCGUAUUUUUUGCAAUUCGAAAUAAUAUUGCUCAUACGCAUGGUUGGCAGAACCGAAAUU -----........(((.((.(((((.(.(((((((.(((..(((((.....)))))......))).))))))).).))))))).)))....... ( -25.50, z-score = -1.62, R) >droSim1.chrX_random 2621009 78 - 5698898 -----CACGA---GUUGUGGCCAACGCGGCGUAUGAGUAUUUUUUACGAUUCGAAAUAAUAUUGCUCAUACGCACGGUUGGCAGAA-------- -----(((..---...)))((((((.(((((((((((((.......................))))))))))).))))))))....-------- ( -30.30, z-score = -4.43, R) >droSec1.super_34 276927 85 - 457129 -----CACGAGUUGUUGUGGCCAACGCGGCGUAUGAGUAUUUUUUACGAUUCGAAAUAAUAUUGCUCAUACGCACAGUUGGCAGAACCAA---- -----(((((....)))))((((((...(((((((((((.......................)))))))))))...))))))........---- ( -30.90, z-score = -4.17, R) >consensus _____CACGAGUUGUUGUGGCCAACGCGGCGUAUGAGUAUUUUUUACGAUUCGAAAUAAUAUUGCUCAUACGCACGGUUGGCAGAACCAA____ ...................((((((...(((((((((((.......................)))))))))))...))))))............ (-25.60 = -25.85 + 0.25)

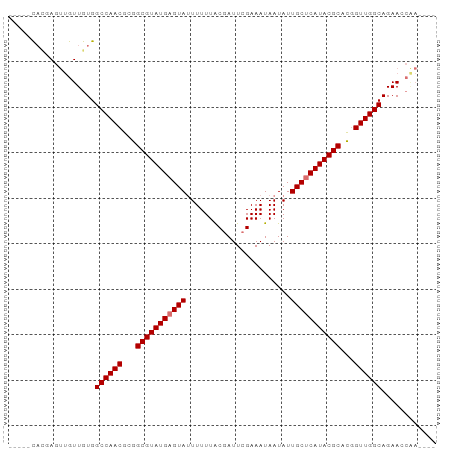
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:27:13 2011