
| Sequence ID | dm3.chrX |
|---|---|
| Location | 8,680,076 – 8,680,152 |
| Length | 76 |
| Max. P | 0.975291 |

| Location | 8,680,076 – 8,680,152 |
|---|---|
| Length | 76 |
| Sequences | 9 |
| Columns | 77 |
| Reading direction | forward |
| Mean pairwise identity | 76.67 |
| Shannon entropy | 0.49051 |
| G+C content | 0.35920 |
| Mean single sequence MFE | -14.63 |
| Consensus MFE | -10.19 |
| Energy contribution | -10.79 |
| Covariance contribution | 0.59 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.70 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.93 |
| SVM RNA-class probability | 0.975291 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chrX 8680076 76 + 22422827 AGAGAAAAUGGCAGCGUUGAAAUAUUUAACGCCGCUAAGUAUGCUACGAAAAUAUCUUUCCCAACAAGCGAACAAA- ........((((.(((((((.....))))))).)))).((.((((..((((.....))))......)))).))...- ( -16.00, z-score = -1.95, R) >droSec1.super_34 54161 76 + 457129 AGAGAAAAUGGCAGCGUUGAAAUAUUUAACGCCGCUAAGUAUGCUACGAAAAUAUCUUUCCCAACAAGCGAACAAA- ........((((.(((((((.....))))))).)))).((.((((..((((.....))))......)))).))...- ( -16.00, z-score = -1.95, R) >droYak2.chrX 9143508 76 - 21770863 AGAGAAAAUGGCAGCGUUGAAAUAUUUAACGCCGCUAAGUAUGCUACGAAAAUAUCGUUCCCAACAAGCGAACAAA- ........((((.(((((((.....))))))).)))).((.((((((((.....))))........)))).))...- ( -17.90, z-score = -2.40, R) >droEre2.scaffold_4690 12328834 75 - 18748788 AGAGAAAAUGGCAGCGUUGAAAUAUUUAACGCCGCUAAGUAUGCUACGAAAAUAUCUUUCCCAA-AAGCGAACAAA- ........((((.(((((((.....))))))).)))).((.((((..((((.....))))....-.)))).))...- ( -16.60, z-score = -2.22, R) >droAna3.scaffold_13047 533520 77 + 1816235 AGGAAAAAUGGCAGCGUUGAAAUAUUUAACGCCGUUAAGUAUGCUACGAAAAUAUUUUGCCAGCAGCAGCGAAAAUA ........((((((((((((.....)))))))(((..((....))))).........)))))((....))....... ( -16.40, z-score = -0.71, R) >dp4.chrXL_group1e 7952624 64 + 12523060 AAAAAAAAUGGCAGCGUUGAAAUAUUUAACGCCGCUAAGUAUGCUACGAAAAUAUUUUGCCGCA------------- ........((((.(((((((.....))))))).))))....(((..((((.....))))..)))------------- ( -15.00, z-score = -1.57, R) >droPer1.super_13 1680651 64 - 2293547 AAAAAAAAUGGCAGCGUUGAAAUAUUUAACGCCGCUAAGUAUGCUACGACAAUAUUUUGCCGCA------------- .........(((((((((((.....))))))).((.......)).............))))...------------- ( -14.30, z-score = -1.19, R) >droWil1.scaffold_181150 3800373 65 - 4952429 AAAAAAAAUGGCACUGUUGAAAU----------GCUAAGUAUGCUACGAAAAUAUU-UUCAGAACAUGCGAAAAAA- ........(((((.(.....).)----------)))).(((((....((((....)-)))....))))).......- ( -8.50, z-score = -0.49, R) >droGri2.scaffold_14853 8653945 57 + 10151454 AACAAAAAUGGCAGCGUUGAAAUAUUUAACGUCGCUAAGUAUACCACAAAAAUAUUU-------------------- ........((((.(((((((.....))))))).))))....................-------------------- ( -11.00, z-score = -2.23, R) >consensus AGAAAAAAUGGCAGCGUUGAAAUAUUUAACGCCGCUAAGUAUGCUACGAAAAUAUUUUUCCCAA_AAGCGAACAAA_ ........((((.(((((((.....))))))).))))........................................ (-10.19 = -10.79 + 0.59)

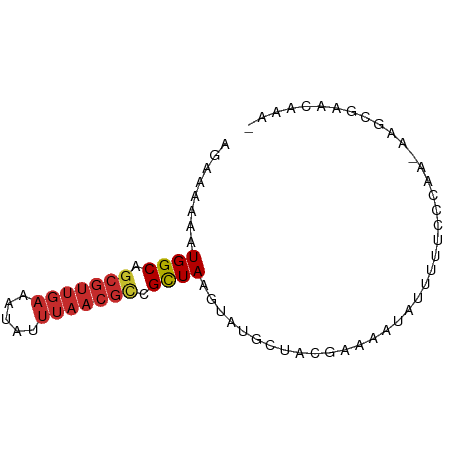
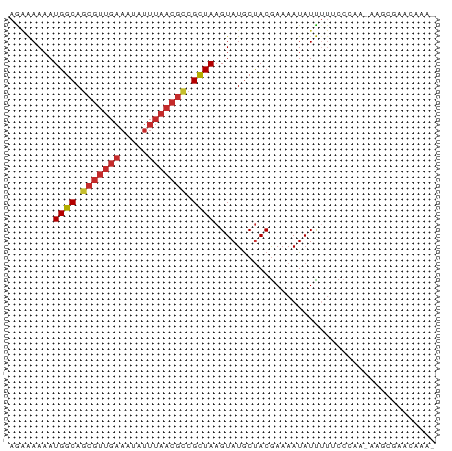
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:26:33 2011