
| Sequence ID | dm3.chrX |
|---|---|
| Location | 8,666,420 – 8,666,479 |
| Length | 59 |
| Max. P | 0.584280 |

| Location | 8,666,420 – 8,666,479 |
|---|---|
| Length | 59 |
| Sequences | 11 |
| Columns | 59 |
| Reading direction | forward |
| Mean pairwise identity | 95.59 |
| Shannon entropy | 0.10149 |
| G+C content | 0.38306 |
| Mean single sequence MFE | -12.07 |
| Consensus MFE | -11.68 |
| Energy contribution | -11.68 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -0.93 |
| Structure conservation index | 0.97 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.584280 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 8666420 59 + 22422827 UUGCUGGCUGUAAAAUUAGGUUAAUUCCUGUCAGUGCGUUGCUGAUUUGAAUUGUCACA ((((.....)))).....(..((((((..((((((.....))))))..))))))...). ( -12.50, z-score = -0.81, R) >droPer1.super_13 1666763 59 - 2293547 UUGUUGGCUGUAAAAUUAGGUUAAUUCCUGUCAGUGCGUUGCUGAUUUGAAUUGUCACA .......(((......)))(.((((((..((((((.....))))))..)))))).)... ( -11.60, z-score = -0.80, R) >dp4.chrXL_group1e 7938667 59 + 12523060 UUGUUGGCUGUAAAAUUAGGUUAAUUCCUGUCAGUGCGUUGCUGAUUUGAAUUGUCACA .......(((......)))(.((((((..((((((.....))))))..)))))).)... ( -11.60, z-score = -0.80, R) >droEre2.scaffold_4690 12315824 59 - 18748788 UUGCUGGCUGUAAAAUUAGGUUAAUUCCUGUCAGUGCGUUGCUGAUUUGAAUUGUCACA ((((.....)))).....(..((((((..((((((.....))))))..))))))...). ( -12.50, z-score = -0.81, R) >droYak2.chrX 9129786 59 - 21770863 UUGCUGGCUGUAAAAUUAGGUUAAUUCCUGUCAGUGAGUUGCUGAUUUGAAUUGUCACA ((((.....)))).....(..((((((..((((((.....))))))..))))))...). ( -12.50, z-score = -0.96, R) >droSec1.super_34 40668 59 + 457129 UUGCUGGCUGUAAAAUUAGGUUAAUUCCUGUCAGUGCGUUGCUGAUUUGAAUUGUCACA ((((.....)))).....(..((((((..((((((.....))))))..))))))...). ( -12.50, z-score = -0.81, R) >droWil1.scaffold_181150 1165782 59 + 4952429 UUGCUGACUGUAAAAUUAGGUUAAUUCCUGUCAGUGCGUUGCUGAUUUGAAUUGUCACA ((((.....)))).....(..((((((..((((((.....))))))..))))))...). ( -12.50, z-score = -1.20, R) >droAna3.scaffold_13047 522053 53 + 1816235 -----GACUGUAAAAUUAGGUUAAUUCCUGUCAGUGCGUCGCUGAUUCGAAUUGUCGC- -----..(((......)))(.((((((..(((((((...)))))))..)))))).)..- ( -12.30, z-score = -1.65, R) >droVir3.scaffold_12726 2483763 59 + 2840439 UUGUUGGCUGUAAAAUUAGGUUAAUUCCUGUCAGUGCGUUGCUGAUUUGAAUUGUCACA .......(((......)))(.((((((..((((((.....))))))..)))))).)... ( -11.60, z-score = -0.80, R) >droMoj3.scaffold_6473 10927649 59 - 16943266 UUGUUGGCUGUAAAAUUAGGUUAAUUCCUGUCAGUGCGUUGCUGAUUUGAAUUGUCACA .......(((......)))(.((((((..((((((.....))))))..)))))).)... ( -11.60, z-score = -0.80, R) >droGri2.scaffold_14853 8636777 59 + 10151454 UUGUUGGCUGUAAAAUUAGGUUAAUUCCUGUCAGUGCGUUGCUGAUUUGAAUUGUCACA .......(((......)))(.((((((..((((((.....))))))..)))))).)... ( -11.60, z-score = -0.80, R) >consensus UUGCUGGCUGUAAAAUUAGGUUAAUUCCUGUCAGUGCGUUGCUGAUUUGAAUUGUCACA .......(((......)))(.((((((..((((((.....))))))..)))))).)... (-11.68 = -11.68 + 0.00)

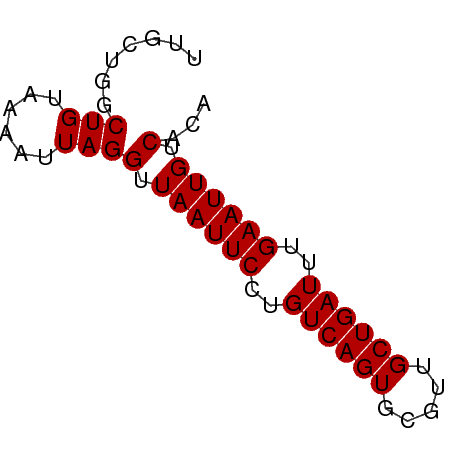
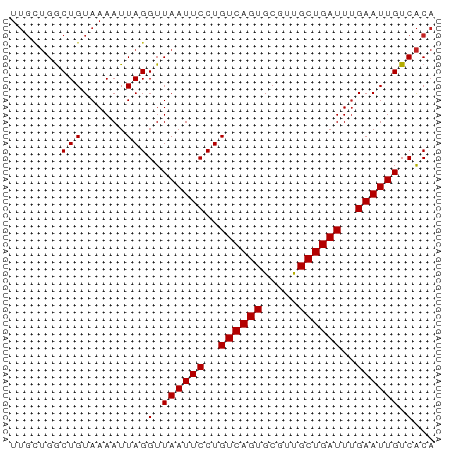
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:26:31 2011