
| Sequence ID | dm3.chrX |
|---|---|
| Location | 8,622,992 – 8,623,103 |
| Length | 111 |
| Max. P | 0.721666 |

| Location | 8,622,992 – 8,623,103 |
|---|---|
| Length | 111 |
| Sequences | 7 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.67 |
| Shannon entropy | 0.38228 |
| G+C content | 0.47546 |
| Mean single sequence MFE | -32.88 |
| Consensus MFE | -17.41 |
| Energy contribution | -18.66 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.721666 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
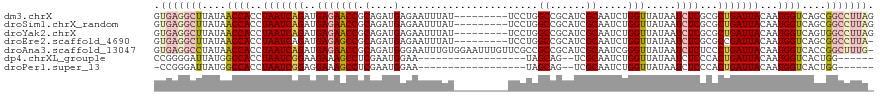
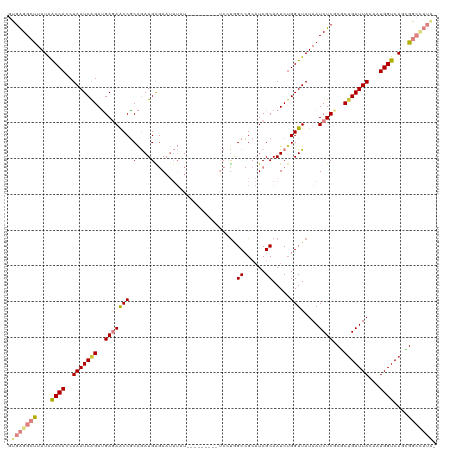
>dm3.chrX 8622992 111 + 22422827 GUGAGGCUUAUAACCACCUAAUCAGAUGAGAACCGCAGAUGAGAAUUUAU---------UCCUGGCCGCAUCGCAAUCUGGUUAUAAUCUCGCGCUGAUUACAAUGGUCAGCGGCCUUAG .(((((((((((((((........((((....(((..(((((....))))---------)..)))...))))......))))))))).....((((((((.....)))))))))))))). ( -34.34, z-score = -2.35, R) >droSim1.chrX_random 706619 111 + 5698898 GUGAGGCUUAUAACCACCUAAUCAGAUGAGAACCGCAGAUGAGAAUUUAU---------UCCUGGCCGCAUCGCAAUCUGGUUAUAAUCUCGCGCUGAUUACAAUGGUCAGCGGCCUUAG .(((((((((((((((........((((....(((..(((((....))))---------)..)))...))))......))))))))).....((((((((.....)))))))))))))). ( -34.34, z-score = -2.35, R) >droYak2.chrX 9082251 111 - 21770863 GUGAGGCUUAUAACCACCUAAUCAGAUGAGAACCGCAGAUGAGAAUUUAU---------UCCUGGCCGCAUCGCAAUCUGGUUAUAAUCUCGCGCUGAUUACAAUGGUCAGUGGCCUUAG .(((((((((((((((........((((....(((..(((((....))))---------)..)))...))))......))))))))).....((((((((.....)))))))))))))). ( -32.44, z-score = -1.85, R) >droEre2.scaffold_4690 12272847 110 - 18748788 GUGAGGCUUAUAACCACCUAAUCAGAUGAGAGCCGCAGAUGAGAAUUUAU---------UCCUGGCCGCAUCGCAAUCUGGUUAUAAUCUCGCGCCGAUUACAAUGGUCAGCGGCCUUA- ((((((.(((((((((........((((.(.((((..(((((....))))---------)..))))).))))......)))))))))))))))((((...((....))...))))....- ( -35.64, z-score = -2.19, R) >droAna3.scaffold_13047 480351 119 + 1816235 GUGAGGCCUAUAACCACCUAAUCAGAUGAGAACCGCAGAUGGGAAUUUGUGGAAUUUGUUCGCCGCCGCAUCGCAAUCGGGUUAUAAUCUCUCCCUGAUUACAAUGGUCACCGGCUUUG- ..((((((....((((..(((((((..((((.((((((((....))))))))...((((...(((..((...))...)))...))))))))...)))))))...))))....)))))).- ( -35.60, z-score = -1.41, R) >dp4.chrXL_group1e 7892453 94 + 12523060 CCGGGGAUUAUGGCCACCUAAUCGGAAGAAAGCCUCGAAUGGAA------------------UAGCAG--UCGCAAUCUGGUUAUAAUCUCCCACUGAUUACAAUGGUCACUGG------ ..((((((((((((((.....((....))...((......))..------------------..((..--..))....))))))))))))))((.(((((.....))))).)).------ ( -27.60, z-score = -1.47, R) >droPer1.super_13 1619230 93 - 2293547 -CCGGGAUUAUGGCCACCUAAUCGGAGGAAAGCCUCGAAUGGAA------------------UAGCAG--UCGCAAUCUGGUUAUAAUCUCCCACUGAUUACAAUGGUCACUGG------ -..(((((((((((((.(((.(((.(((....)))))).)))..------------------..((..--..))....)))))))))))))(((.(((((.....))))).)))------ ( -30.20, z-score = -2.24, R) >consensus GUGAGGCUUAUAACCACCUAAUCAGAUGAGAACCGCAGAUGAGAAUUUAU_________UCCUGGCCGCAUCGCAAUCUGGUUAUAAUCUCGCGCUGAUUACAAUGGUCAGCGGCCUUA_ .(((((((....((((..(((((((..(((((((.(....).......................((......)).....))).....))))...)))))))...))))....))))))). (-17.41 = -18.66 + 1.25)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:26:25 2011