
| Sequence ID | dm3.chrX |
|---|---|
| Location | 8,561,173 – 8,561,270 |
| Length | 97 |
| Max. P | 0.692754 |

| Location | 8,561,173 – 8,561,268 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 69.57 |
| Shannon entropy | 0.49820 |
| G+C content | 0.62932 |
| Mean single sequence MFE | -34.07 |
| Consensus MFE | -18.39 |
| Energy contribution | -18.82 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.31 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.692754 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 8561173 95 + 22422827 GGAAAAGGGGCAGAGUUGCGGGUUAAUCAGUGGCGCGCCUACAAGUGGCGGGCUACCGAAAAGCCCCCUUUUCCUCGCCCCCCCCCCUCCAAAUC (((...(((((.(((..(.(((((..((.(((((.((((.......)))).))))).))..))))).)..)))...)))))......)))..... ( -40.30, z-score = -2.23, R) >droAna3.scaffold_13248 797868 89 - 4840945 AGCGAAGGUUUUGGGCUUGGGGUCAGUCGGUGCCGCGCCUACAAAUGAUCGCCCCCCCUCCACCCCACCUGCCAUCGCACUUCUCCACC------ .((((.(((..((((...(((((((((.((((...)))).))...)))......)))).....))))...))).))))...........------ ( -26.30, z-score = 0.04, R) >droEre2.scaffold_4690 12195667 84 - 18748788 GAAAAGGGGGCAGGGUUGCGGGUCAAUCAGUGCCGCGCCUACAAGUGGCGGGCUACCGAAAAGCCCCCUUUUCCUCACCCGCCC----------- (((((((((((..(((((.....))))).(((((.((((.......))))))).))......)))))))))))...........----------- ( -37.30, z-score = -1.52, R) >droYak2.chrX 9019830 93 - 21770863 -GAAAAGGGGCAGAGUUGCGGGUCAAUCAGUGCCGCGCCUACAAGUGGCGGGCUACCGAAAAGCCCCCUUUUCCUCACCCAACCCCACCCCAUC- -.....((((..(.((((.((..........(((((........)))))(((((.......)))))...........)))))).)..))))...- ( -32.40, z-score = -1.55, R) >consensus GGAAAAGGGGCAGAGUUGCGGGUCAAUCAGUGCCGCGCCUACAAGUGGCGGGCUACCGAAAAGCCCCCUUUUCCUCACCCACCCCCACC______ ......((((.((((....((((...((.(((.(.((((.......)))).).))).))...)))).)))).))))................... (-18.39 = -18.82 + 0.44)


| Location | 8,561,175 – 8,561,270 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 70.46 |
| Shannon entropy | 0.48310 |
| G+C content | 0.63779 |
| Mean single sequence MFE | -33.52 |
| Consensus MFE | -18.39 |
| Energy contribution | -18.82 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.29 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.641002 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 8561175 95 + 22422827 AAAAGGGGCAGAGUUGCGGGUUAAUCAGUGGCGCGCCUACAAGUGGCGGGCUACCGAAAAGCCCCCUUUUCCUCGCCCCCCCCCCUCCAAAUCCC ....(((((.(((..(.(((((..((.(((((.((((.......)))).))))).))..))))).)..)))...)))))................ ( -38.50, z-score = -2.32, R) >droAna3.scaffold_13248 797870 89 - 4840945 CGAAGGUUUUGGGCUUGGGGUCAGUCGGUGCCGCGCCUACAAAUGAUCGCCCCCCCUCCACCCCACCUGCCAUCGCACUUCUCCACCCC------ .((((((....(((.((((((.((..((.((..((........))...)).))..))..))))))...)))...)).))))........------ ( -23.50, z-score = 0.67, R) >droEre2.scaffold_4690 12195669 84 - 18748788 AAAGGGGGCAGGGUUGCGGGUCAAUCAGUGCCGCGCCUACAAGUGGCGGGCUACCGAAAAGCCCCCUUUUCCUCACCCGCCCCC----------- ...((((((.((((.(.(((.........(((((........)))))(((((.......))))).....))).)))))))))))----------- ( -39.70, z-score = -1.97, R) >droYak2.chrX 9019832 93 - 21770863 -AAAGGGGCAGAGUUGCGGGUCAAUCAGUGCCGCGCCUACAAGUGGCGGGCUACCGAAAAGCCCCCUUUUCCUCACCCAACCCCACCCCAUCCC- -...((((..(.((((.((..........(((((........)))))(((((.......)))))...........)))))).)..)))).....- ( -32.40, z-score = -1.54, R) >consensus AAAAGGGGCAGAGUUGCGGGUCAAUCAGUGCCGCGCCUACAAGUGGCGGGCUACCGAAAAGCCCCCUUUUCCUCACCCACCCCCACCCC______ ....((((.((((....((((...((.(((.(.((((.......)))).).))).))...)))).)))).))))..................... (-18.39 = -18.82 + 0.44)
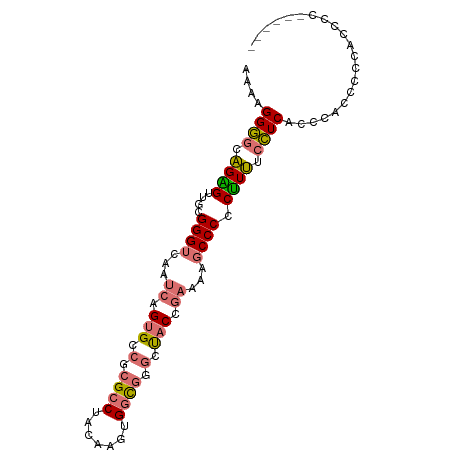


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:26:16 2011