
| Sequence ID | dm3.chrX |
|---|---|
| Location | 8,293,393 – 8,293,446 |
| Length | 53 |
| Max. P | 0.992665 |

| Location | 8,293,393 – 8,293,446 |
|---|---|
| Length | 53 |
| Sequences | 12 |
| Columns | 53 |
| Reading direction | forward |
| Mean pairwise identity | 96.51 |
| Shannon entropy | 0.07253 |
| G+C content | 0.48428 |
| Mean single sequence MFE | -16.69 |
| Consensus MFE | -14.87 |
| Energy contribution | -14.61 |
| Covariance contribution | -0.26 |
| Combinations/Pair | 1.17 |
| Mean z-score | -3.22 |
| Structure conservation index | 0.89 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.56 |
| SVM RNA-class probability | 0.992665 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
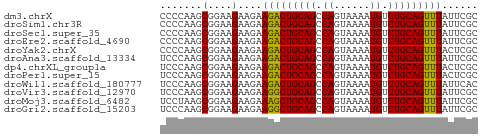
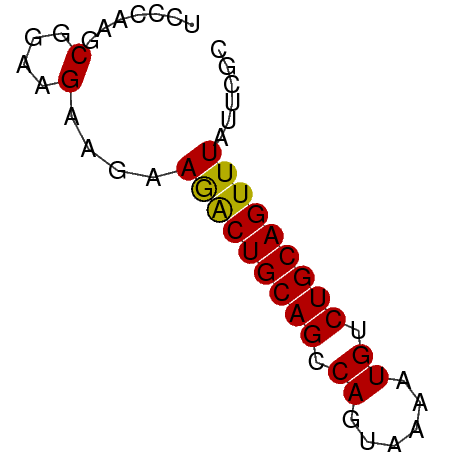
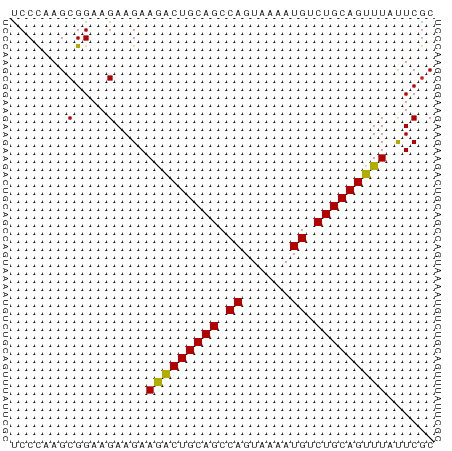
>dm3.chrX 8293393 53 + 22422827 CCCCAAGCGGAAGAAGAAGACUGCAGCCAGUAAAAUGUCUGCAGUUUAUUCGC ..((....))..(((..(((((((((.((......)).))))))))).))).. ( -16.60, z-score = -3.56, R) >droSim1.chr3R 5820725 53 + 27517382 CCCCAAGCGGAAGAAGAAGACUGCAGCCAGUAAAAUGUCUGCAGUUUAUUCGC ..((....))..(((..(((((((((.((......)).))))))))).))).. ( -16.60, z-score = -3.56, R) >droSec1.super_35 143030 53 + 471130 CCCCAAGCGGAAGAAGAAGACUGCAGCCAGUAAAAUGUCUGCAGUUUAUUCGC ..((....))..(((..(((((((((.((......)).))))))))).))).. ( -16.60, z-score = -3.56, R) >droEre2.scaffold_4690 10658391 53 + 18748788 CCCCAAGCGGAAGAAGAAGACUGCAGCCAGUAAAAUGUCUGCAGUUUAUUCGC ..((....))..(((..(((((((((.((......)).))))))))).))).. ( -16.60, z-score = -3.56, R) >droYak2.chrX 8750084 53 - 21770863 CCCCAAGCGGAAGAAGAAGACUGCAGCCAGUAAAAUGUCUGCAGUUUACUCGC ......(((.....((.(((((((((.((......)).))))))))).))))) ( -15.80, z-score = -2.72, R) >droAna3.scaffold_13334 542042 53 - 1562580 UCCCAAGCGGAAGAAGAAGACUGCAGCCAGUAAAAUGUCUGCAGUUUAUUCGC (((.....))).(((..(((((((((.((......)).))))))))).))).. ( -17.30, z-score = -3.67, R) >dp4.chrXL_group1a 117760 53 - 9151740 UCCCAAGCGGAAGAAGAAGACUGCAGCCAGUAAAAUGUCUGCAGUUUACUCGC (((.....))).((.(.(((((((((.((......)).))))))))).))).. ( -16.10, z-score = -2.69, R) >droPer1.super_15 755150 53 - 2181545 UCCCAAGCGGAAGAAGAAGACUGCAGCCAGUAAAAUGUCUGCAGUUUACUCGC (((.....))).((.(.(((((((((.((......)).))))))))).))).. ( -16.10, z-score = -2.69, R) >droWil1.scaffold_180777 4588517 53 - 4753960 UCCCAAGCGGAAGAAGAAGACUGCAGCCAGUAAAAUGUCUGCAGUUUAUUCAC (((.....))).(((..(((((((((.((......)).))))))))).))).. ( -17.30, z-score = -4.06, R) >droVir3.scaffold_12970 7963844 53 + 11907090 UCCCAAGCGGAAGAAGAAGGCUGCAGCCAGUAAAAUGUCUGCAGUUUAUUCGC (((.....))).(((..(((((((((.((......)).))))))))).))).. ( -16.80, z-score = -2.64, R) >droMoj3.scaffold_6482 1167336 53 - 2735782 UCCUAAGCGGAAGAAGAAAGCUGCAGCCAGUAAAAUGUCUGCAGUUUAUUCGC (((.....))).(((..(((((((((.((......)).))))))))).))).. ( -17.70, z-score = -3.33, R) >droGri2.scaffold_15203 892302 53 - 11997470 UCCCAAGCGGAAGAAGAAGGCUGCAGCCAGUAAAAUGUCUGCAGUUUAUUCGC (((.....))).(((..(((((((((.((......)).))))))))).))).. ( -16.80, z-score = -2.64, R) >consensus UCCCAAGCGGAAGAAGAAGACUGCAGCCAGUAAAAUGUCUGCAGUUUAUUCGC .......(....)....(((((((((.((......)).)))))))))...... (-14.87 = -14.61 + -0.26)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:25:42 2011