
| Sequence ID | dm3.chrX |
|---|---|
| Location | 8,286,543 – 8,286,643 |
| Length | 100 |
| Max. P | 0.564418 |

| Location | 8,286,543 – 8,286,643 |
|---|---|
| Length | 100 |
| Sequences | 7 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 68.15 |
| Shannon entropy | 0.61101 |
| G+C content | 0.49828 |
| Mean single sequence MFE | -20.28 |
| Consensus MFE | -8.25 |
| Energy contribution | -8.19 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.24 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.564418 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chrX 8286543 100 - 22422827 CAUUCCCUUGUCAGGUAUUUCAGCUUGUUGUUUGCUGC-------AGUUAUGUGUUUAACUGCCACACACAUA-CACCGAGCACCUCCCCCCCCCCCCCCCCUCUUUA- .....(((....))).......(((((.(((.((..((-------(((((......)))))))......)).)-)).)))))..........................- ( -18.10, z-score = -2.56, R) >droSim1.chrX 6593629 101 - 17042790 CAUUCCCUUGUCAGGUAUUUCAGCUUGUUGUUUGCUGC-------AGUUAUGUGUUUAACUGCCACACACACACACACACCCCCCUAGCACCUCCUCCGCCCCCCUCA- ........((..(((..........((((((.((..((-------(((((......)))))))..)).))).)))........)))..))..................- ( -16.47, z-score = -1.04, R) >droSec1.super_35 135950 99 - 471130 CAUUCCCUUGUCAGGUAUUUCAGCUUGUUGUUUGCUGC-------AGUUAUGUGUUUAACUGC--CACACACACACACACCCCCCUAGCACCUCCCCCGCCCCCUUCA- ........((..(((..........((((((.((..((-------(((((......)))))))--..)).))).)))......)))..))..................- ( -15.59, z-score = -0.91, R) >droEre2.scaffold_4690 10651116 96 - 18748788 CAUUCCCUUGUCAGGUAUUUCAGCUUGUUGUUUGCUGC-------AGUUAUGUGUUUAACUGCCACACACAGG-ACACACACACACACCCCCCCGCCCCCGCCU----- ...(((.....(((((......))))).(((.((..((-------(((((......)))))))..)).)))))-).............................----- ( -17.10, z-score = 0.13, R) >droAna3.scaffold_13248 705334 101 - 4840945 CAUUCCCUUGUCAGGUAUUUCAGCUUGUUGUUUGCUGC-------AGUUAUGUGUUUAACUGCCACACACACAUACCC-CUUAGGUAUCUAGGUAUCGUCCCCCGUUGG .....(((....))).....((((.(((.((.((..((-------(((((......)))))))..)).))))).....-....(((((....))))).......)))). ( -19.90, z-score = -0.01, R) >droPer1.super_18 1262239 89 + 1952607 CAUUGCCCCGUCAGGUAUGCCAGCUUGUUGUGUGUUGUGUGUUGGAGUUAUGUGUGUAACUGCCAUACACACAAUGUCCUUCUUGCUUU-------------------- ...((((......))))....(((..(..(..(((((((((((((((((((....)))))).))).))))))))))..)..)..)))..-------------------- ( -27.40, z-score = -2.15, R) >dp4.chrXL_group1e 10682779 89 + 12523060 CAUUGCCCCGUCAGGUAUGCCAGCUUGUUGUGUGUUGUGUGUUGGAGUUAUGUGUGUAACUGCCAUACACACAAUGUCCUUCUUGCUUU-------------------- ...((((......))))....(((..(..(..(((((((((((((((((((....)))))).))).))))))))))..)..)..)))..-------------------- ( -27.40, z-score = -2.15, R) >consensus CAUUCCCUUGUCAGGUAUUUCAGCUUGUUGUUUGCUGC_______AGUUAUGUGUUUAACUGCCACACACACA_ACACACUCACCUAUC_CC_CCCCC_CCCCC_U___ .............(((....((((.........))))........(((((......))))))))............................................. ( -8.25 = -8.19 + -0.06)


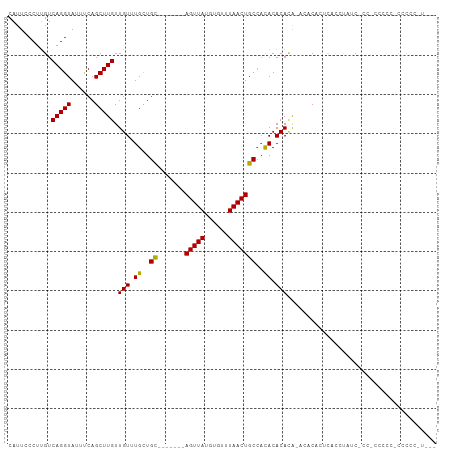
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:25:41 2011