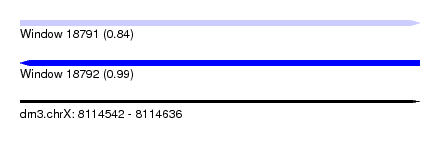

| Sequence ID | dm3.chrX |
|---|---|
| Location | 8,114,542 – 8,114,636 |
| Length | 94 |
| Max. P | 0.989553 |

| Location | 8,114,542 – 8,114,636 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 85.69 |
| Shannon entropy | 0.26789 |
| G+C content | 0.46790 |
| Mean single sequence MFE | -24.64 |
| Consensus MFE | -19.51 |
| Energy contribution | -19.90 |
| Covariance contribution | 0.39 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.85 |
| SVM RNA-class probability | 0.835723 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
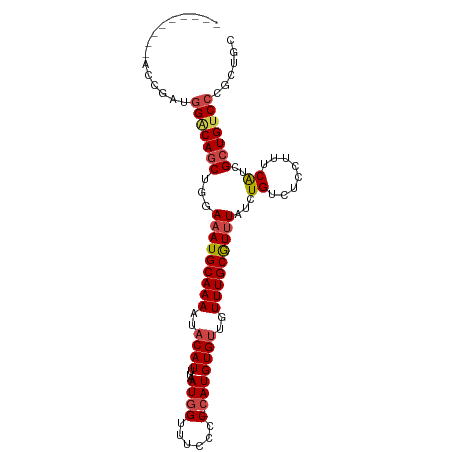
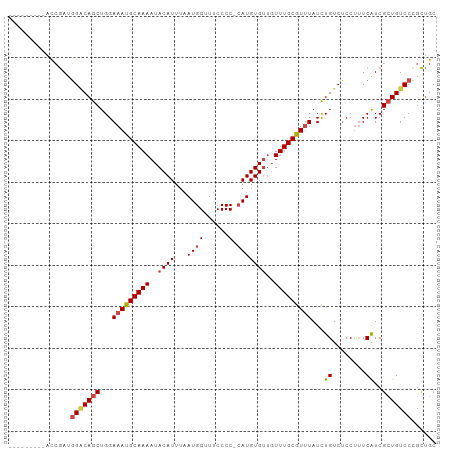
>dm3.chrX 8114542 94 + 22422827 ---------ACCGAUGGGCAGCUGGAAAUGCAAAAUACAUUUAAUGGUUUCCCCACAUGUGUUGUUUGCGUUUAUCUGUCUCCUUUCAUCGCUGUCCCGCUGC ---------......(((((((...(((((((((..((((....(((.....)))...))))..)))))))))...((........))..)))))))...... ( -23.90, z-score = -1.29, R) >droSim1.chrX 6469941 93 + 17042790 ---------ACCGAUGGACAGCUGGAAAUGCAAAAUACAUUUAAUGGUUUCCCC-CAUGUGUUGUUUGCGUUUAUCUGUCUCCUUUCAUCGCUGUCCCGCUGC ---------......(((((((...(((((((((..((((...((((......)-)))))))..)))))))))...((........))..)))))))...... ( -25.00, z-score = -2.57, R) >droSec1.super_44 205691 93 + 251150 ---------ACCGAUGGACAGCUGGAAAUGCAAAAUACAUUUAAUGGUUUCCCC-CAUGUGUUGUUUGCGUUUAUCUGUCUCCUUUCAUCGCUGUCCCGCUGC ---------......(((((((...(((((((((..((((...((((......)-)))))))..)))))))))...((........))..)))))))...... ( -25.00, z-score = -2.57, R) >droYak2.chrX 8570909 102 - 21770863 UAUGCACAUACCGAUGGGCAGCUGGAAAUGCAAAAUACAUUUAAUGGUUUCCCC-CAUGUGUUGUUUGCGUUUAUCUGUCUCCUUUCAUCGCUGUCCCGCUGC ...((..(((.((((((((((....(((((((((..((((...((((......)-)))))))..)))))))))..)))))......))))).)))...))... ( -25.10, z-score = -0.94, R) >droEre2.scaffold_4690 10481911 92 + 18748788 ---------ACCGAUGGCCAGCUGGAAAUGCAAAAUACAUUGAAUGGUUUCCCC--AUGUGUUGUUUGCGUGUAUCUGUCUCCUUUCGUCGCUGUCCAGCUGC ---------.........((((((((...(((((..((((...((((.....))--))))))..)))))(((.................)))..)))))))). ( -23.83, z-score = -1.20, R) >droAna3.scaffold_13417 463660 99 + 6960332 GAGAGACAGUUGAACCGACAGCUGGAAAUGCAAAAUACAUUUAAUGGCUUCCCC-CAUGUG---UUUGCAUUUAUUUGUCGCCCCUCAUCGAUGUCCUGUCGC ((.(((((..(((..((((((....(((((((((.((((.....(((......)-))))))---)))))))))..))))))....)))....))).)).)).. ( -25.00, z-score = -1.84, R) >consensus _________ACCGAUGGACAGCUGGAAAUGCAAAAUACAUUUAAUGGUUUCCCC_CAUGUGUUGUUUGCGUUUAUCUGUCUCCUUUCAUCGCUGUCCCGCUGC ...............(((((((...(((((((((..((((.....((....)).....))))..)))))))))...((........))..)))))))...... (-19.51 = -19.90 + 0.39)

| Location | 8,114,542 – 8,114,636 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 85.69 |
| Shannon entropy | 0.26789 |
| G+C content | 0.46790 |
| Mean single sequence MFE | -28.38 |
| Consensus MFE | -22.29 |
| Energy contribution | -22.32 |
| Covariance contribution | 0.03 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.71 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.37 |
| SVM RNA-class probability | 0.989553 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
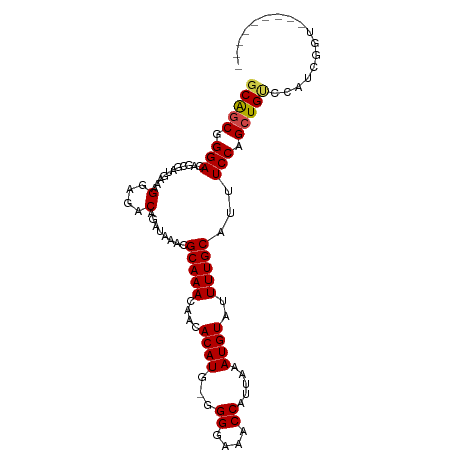
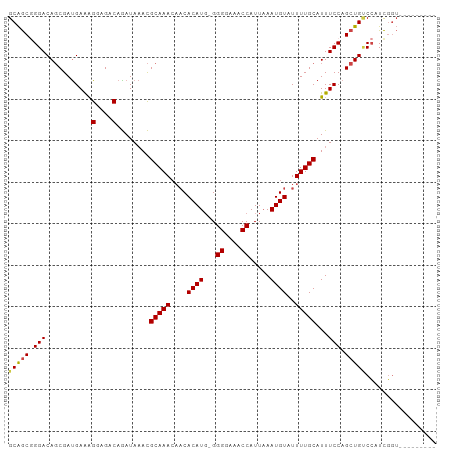
>dm3.chrX 8114542 94 - 22422827 GCAGCGGGACAGCGAUGAAAGGAGACAGAUAAACGCAAACAACACAUGUGGGGAAACCAUUAAAUGUAUUUUGCAUUUCCAGCUGCCCAUCGGU--------- .....(((.((((...(((((....)........(((((....((((...((....)).....))))..))))).))))..)))))))......--------- ( -28.10, z-score = -2.61, R) >droSim1.chrX 6469941 93 - 17042790 GCAGCGGGACAGCGAUGAAAGGAGACAGAUAAACGCAAACAACACAUG-GGGGAAACCAUUAAAUGUAUUUUGCAUUUCCAGCUGUCCAUCGGU--------- ((....(((((((...(((((....)........(((((....((((.-.((....)).....))))..))))).))))..)))))))....))--------- ( -29.30, z-score = -3.62, R) >droSec1.super_44 205691 93 - 251150 GCAGCGGGACAGCGAUGAAAGGAGACAGAUAAACGCAAACAACACAUG-GGGGAAACCAUUAAAUGUAUUUUGCAUUUCCAGCUGUCCAUCGGU--------- ((....(((((((...(((((....)........(((((....((((.-.((....)).....))))..))))).))))..)))))))....))--------- ( -29.30, z-score = -3.62, R) >droYak2.chrX 8570909 102 + 21770863 GCAGCGGGACAGCGAUGAAAGGAGACAGAUAAACGCAAACAACACAUG-GGGGAAACCAUUAAAUGUAUUUUGCAUUUCCAGCUGCCCAUCGGUAUGUGCAUA .....(((.((((...(((((....)........(((((....((((.-.((....)).....))))..))))).))))..)))))))............... ( -27.70, z-score = -1.53, R) >droEre2.scaffold_4690 10481911 92 - 18748788 GCAGCUGGACAGCGACGAAAGGAGACAGAUACACGCAAACAACACAU--GGGGAAACCAUUCAAUGUAUUUUGCAUUUCCAGCUGGCCAUCGGU--------- .((((((((..((..(....)..).)........(((((....((((--.((....)).....))))..)))))...)))))))).........--------- ( -28.30, z-score = -3.21, R) >droAna3.scaffold_13417 463660 99 - 6960332 GCGACAGGACAUCGAUGAGGGGCGACAAAUAAAUGCAAA---CACAUG-GGGGAAGCCAUUAAAUGUAUUUUGCAUUUCCAGCUGUCGGUUCAACUGUCUCUC ..(((((..((....)).(((.(((((...(((((((((---.((((.-.((....)).....))))..))))))))).....))))).)))..))))).... ( -27.60, z-score = -1.69, R) >consensus GCAGCGGGACAGCGAUGAAAGGAGACAGAUAAACGCAAACAACACAUG_GGGGAAACCAUUAAAUGUAUUUUGCAUUUCCAGCUGUCCAUCGGU_________ (((((.(((...........(....)........(((((....((((...((....)).....))))..)))))...))).)))))................. (-22.29 = -22.32 + 0.03)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 01:25:22 2011